From farmers to bioengineers
Sowing genes, harvesting molecules

Twenty-first-century agriculture faces major challenges that urgently need to be answered. In the last decade, new breeding technologies have been developed that can help meet these challenges. These technologies are not only more accurate and efficient, but are also simpler and more accessible, which will facilitate the progressive democratisation of agricultural biotechnology. In this text we discuss future agricultural development in terms of technological democratisation and regulatory relaxation. In this scenario one would expect an increase in the diversity of cultivated varieties and species, the strong development of biofactory crops and, in the long term, the emergence of increasingly fit «smart» crops.
Keywords: new plant breeding technologies, genetic editing, synthetic biology, biofactory plants.
Introduction
The citizens of developed countries tend to consider their food supply as guaranteed at our current technological levels and thus they underestimate the strategic importance of agricultural biotechnology. However, this could not be further from reality. According to the latest report on the future of agriculture published by the Food and Agriculture Organization of the United Nations (FAO, 2018), the world population will reach 10 billion in 2050, leading to an increase in cultivated land and a consequent decline in forested areas. At the same time, climate change is threatening to reduce the productivity of our crops. New plagues are putting our plants at risk, and this is favoured by trade and climatic conditions that encourage the development of such diseases. In turn, the fight against these emerging pests will require increased use of plant protection products, consequently generating potentially toxic by-products. In this context of demographic and environmental pressure, improving our agricultural production systems is essential if we are to achieve a sustainable future (Figure 1).
«Genetic improvement can help to create crops with genetic mechanisms that can allow them to defend themselves from new threats»
One of the most efficient strategies available to address these challenges is plant breeding. This involves providing our crops with genetic mechanisms so they can defend themselves from new threats without the need for external agents. Furthering plant breeding based on sustainability factors such as genetic resistance to pests and diseases, resilience, and adaptation to salinity, droughts, high temperatures, etc., is one of the biggest challenges of future plant biotechnology. However, as has happened in the past with other technological fields, our future agro-biotechnological development model will largely depend on how new plant breeding technologies are perceived by society. A good example is the evolution of computing and telecommunications. Computing started in the last century as a deeply elitist technology monopolised by states and large companies, and this monopoly generated mistrust among the population. All this changed radically with the democratisation brought about by the emergence of the personal computer, which was possible thanks to the decrease in technology prices because of the standardisation of electronic components.

Figure 2. Biotechnology is often considered an elitist technology. However, new breeding technologies are now allowing for a «democratisation» leap similar to that in the field of information technology created by the introduction of personal computers. / HCC Public Information Office
To date, agricultural biotechnology has also been conducted as an elitist technology, made evident by the small number of companies that concentrate most of the world’s seed production (Figure 2). As we will see below, new breeding technologies, together with the vast corpus of knowledge generated in recent decades by disciplines such as genomics and the study of plant physiology, now places us in a technological position that would allow a «democratisation» leap, similar to that created by the introduction of personal computers to the field of information technology. However, meeting the right technological conditions does not ensure that such a transition will take place. It will all depend on how such technology is perceived, accepted and, eventually, incorporated into our societies. This could range from an ultra-conservative scenario that appears to reject new technologies, to an ultra-liberal scenario that dispenses almost entirely with any regulation. It is reasonable to expect that these extreme scenarios will not take place, but even so, small differences in how regulatory barriers are set can yield completely different results. A more restrictive stance based on the rigorous application precautionary principles would create obstacles for technological development that could only be overcome by a few parties with sufficient financial capacity. This would then place biotechnology in the hands of only a few people. Conversely, a scenario with more accessible regulation could stimulate creativity, competition, and encourage the emergence of more actors, including those operating in rural areas, without affecting food or environmental security.
Here, we will examine what the agricultural food landscape of the future might look like, assuming the most favourable scenario of responsible innovation. To do this, we will first look at how some of the new enabling technologies have pushed agricultural breeding towards a tipping point, and then we will imagine how these technologies might, in a context that favours innovation, lead to the plants of the future.
New tools for breeding
Breeding involves modifying some of the genetic instructions contained in a species’ genome to suit our needs. Humans have been modifying crops’ genomes since the Neolithic era, selecting for the random mutations that were most beneficial to us. However, until the last century, our ability to perform genetic editing was limited to the emergence of spontaneous mutations. More recently, marker-assisted breeding has accelerated the incorporation of mutations from the genomes of other, related varieties or species into our crops.
«Furthering genetic improvement is one of the biggest challenges of future plant biotechnology research»
Chemical mutagenesis and radiation techniques have also facilitated the rapid generation of new genetic variability that can be incorporated into breeding programmes. However, it was not until the early 1980s that plant biotechnology tools allowed the transfer of «engineered» (i.e., non-random) instructions into plant genomes for the first time (Barton et al., 1983; Herrera-Estrella et al., 1983). Current genetically modified (transgenic) plants usually include only one or, at most, a few «designer» genes resulting from DNA recombination, i.e., a minuscule amount compared to the nearly 50,000 genes that make up the genome of a plant like the soybean (Schmutz et al., 2010). Nonetheless, the impact of plant transgenesis has been so profound that 75 % of the soybeans planted today carry these few genetic modifications.

Figure 3. An explosion of diversity is expected in the next ten years, both in varieties (thanks to the improvement of local varieties) and in cultivated species. / PxHere
In the context of this first generation of biotechnological tools, our ability to edit genomes has increased dramatically in the last decade thanks to a set of new technologies. Among these, two deserve special mention: DNA synthesis and CRISPR genetic editing. DNA synthesis allows us to write genetic instructions in the laboratory. In recent years our capacity to synthesise DNA has grown almost exponentially so that now we can write increasingly longer DNA fragments at a lower cost (Wang et al., 2018). Although the length of DNA chains that can be synthesised at once using purely chemical methods is not unlimited, so-called modular assembly techniques have recently allowed the creation of ever longer constructs by joining DNA fragments with each other as if they were Lego parts (Engler et al., 2008; Vazquez-Vilar et al., 2018). As a result, our ability to «write» complex instructions that can subsequently be transferred to plants has grown enormously, both in quantity and in precision, while the price and effort required to do so has decreased.
The second major technological breakthrough was CRISPR gene-editing technology. Being able to write genetic instructions is of little use if we cannot insert them correctly into the corresponding page of the genetic instruction book. CRISPR/Cas9 proteins, discovered, among others, by the Spanish researcher Francis Mojica (Lander, 2016; Mojica et al., 2009), are the genomic equivalent of powerful search engines. They can allow us to navigate the immense genomic instruction book and either introduce small modifications or incorporate new information into it with precision. The novelty of CRISPR technology is that it allows us to incorporate new information exactly into the chosen line of the desired genetic code page, in the specific location where an instruction makes sense, and nowhere else. It is important to note that CRISPR gene editing has been proven not only to be a hugely efficient technology, but it is also a very affordable one, thus putting all of its potential within the reach of large and small laboratories alike. Thus, it is easy to conclude that combining the growing capability for DNA synthesis with the power and precision allowed by genomic editing could make breeding the starting point of a technological revolution.
From farmers to bioengineers
Next, we will examine what the plants of the future might look like given these new technological capabilities. As described above, we will assume that future regulatory scenarios will be flexible enough to make their adoption by small and medium-sized companies, or their public counterparts, feasible. The first consequence of such a scenario would be the creation of a new innovation ecosystem associated with agriculture. One of the main problems of the current system is the tendency towards uniformity in the development of elite varieties to replace native ones. The «democratisation» of agrobiotechnology would facilitate the direct introduction of genetic innovations such as resistance, organoleptic, or nutritional characteristics, etc. into local varieties. This would bring genetic design closer to local producers. In other words, the «democratisation» of biotechnology would enable a professional evolution from farmers to «farmer-bioengineers», or even «biohackers» – professionals specialising in different phases of the agricultural production cycle – from designing the variety to producing it in the field. Thanks to this creative environment, the plants of the future would be capable of yielding pleasant surprises. Here are some examples of what this creative new agriculture might look like.
Explosion of cultivated biodiversity
The most immediate change to expect over a ten-year period would be an explosion in the diversity of both varieties and species (Figure 3). First, varietal diversity would be expanded by facilitating the improvement of local varieties. In the medium term, we could also expect an increase in the number of cultivated species. In agriculture we use only a small part of the catalogue of plant species that exist in nature, since just a few can be domesticated by traditional methods. Today, however, we know many of the genetic factors that have previously allowed the domestication of our crop species. Aided by new breeding techniques, we can now undertake the domestication of new wild species. In a recent example, a group of German and Brazilian researchers managed to improve a species of wild tomato with no agronomic value by editing only six of its genes: they increased the number of fruits per plant tenfold, tripled their size, and quintupled their lycopene content (Khan et al., 2019; Zsögön et al., 2018). Therefore, these researchers partially recapitulated much of the centuries-old process of the domestication of the cultivated tomato in just a couple generations.
«Even today, many of the drugs used in the fight against diseases such as cancer or malaria are extracted from plants»
Similar processes are expected to occur in other wild or semi-domesticated species, many of which can provide greater base resilience or better adaptation to hostile environments. New technologies will even allow us to speed up the speciation processes, as Professor Ralf Bock’s group in Germany recently demonstrated by inducing neopolyploidy with a graft (Fuentes et al., 2014). Starting with two tobacco species, Nicotiana tabacum and Nicotiana glauca, these researchers quickly created a new species (Nicotiana tabauca) by transferring complete nuclear genomes from one plant to another. After the transfer, a rapid genomic reorganisation process, similar to that occurring during natural speciation, took place. Thus, it is likely that we will be able to quickly generate new species on which we can test agronomic characteristics, composition, etc., in order to create radically new crops.

Figure 4. The species Nicotiana benthamiana, friendly known as «Benthy», is widely used to produce vaccines, antibodies, and new products to replace antibiotics. Genetic engineering allows us to use plants as biofactories to obtain biomolecules used in pharmaceuticals, medicine, and cosmetics. / CSIC-CRAG
Biofactory crops: «Made in Benthy»
The above example helps us to underline the fact that, in addition to food production, there is a millenary tradition of using plants as factories to obtain useful compounds including fibres, biomaterials, and medicines. Even today, many of the drugs used in the fight against diseases such as cancer or malaria are extracted from plants. Genetic engineering allows us to use plants as biofactories to obtain biomolecules used in pharmaceuticals, medicine, or cosmetics. For example, the Australian species Nicotiana benthamiana (Figure 4), a relative of tobacco (commonly known as Benthy), is widely used to produce vaccines, antibodies, and new products to replace antibiotics. Indeed, the antidote that Western missionaries received in the recent 2014 Ebola crisis was produced from N. benthamiana (Qiu et al., 2014).
More recently, the Canadian company Medicago completed phase III of its human clinical testing of a seasonal flu vaccine which was also «made in Benthy» (Pillet et al., 2016). It is very likely that in five to ten years’ time our vaccines could be produced using plant-based antigens, and our food will be preserved with «made in Benthy» colicins (antibacterial proteins) similar to those recently developed by the German company Icon Genetics (Stephan et al., 2017). All these examples show a new side of agricultural production, which in turn, sets new previously unexplored goals for breeding, such as enhancing the efficiency, quality, and stability of recombinant proteins or crop biosafety, among others. Indeed, the European project Newcotiana, which brings together nineteen European and Australian research groups, aims at improving the biofactory capacity of Benthy plants and their relative, cultivated tobacco. With the implementation of these improvements, it is expected that the land area dedicated to biofactories in the future will increase, displacing other less socially acceptable crops such as smoking tobacco.
«Sexy», «smart», plants with increased abilities
What can we expect beyond twenty-years’ time? What are the limits of plant breeding? Can we provide our crops with new functionalities beyond those we observe in nature? Some might say everything was already invented by evolution, but it appears that humans may be able to provide abilities that biological evolution did not. One example of this is weather forecasting. It is technologically possible to design «smart» plants that can respond to certain weather-based triggers by, for example, producing defensive compounds such as antifreeze proteins when a frost is likely. Similarly, the current state of technology would allow us to design plants that respond to the detection of a pest by producing defensive volatile compounds such as sexual pheromones that confuse insects.
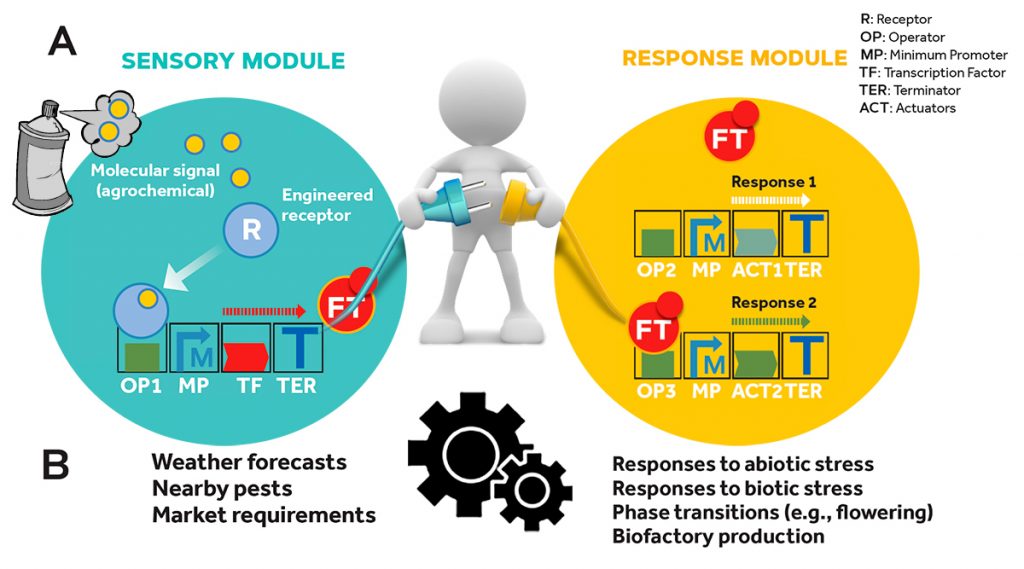
Figure 5. This diagram shows how plants are equipped with new genetic circuits which turn them into «smart» plants. (A) The principles of modularity and standardisation, typical of synthetic biology, facilitate the physical connection between genetic elements and, therefore, the construction of increasingly complex regulatory circuits. (B) Applied to genetic improvement, these circuits would allow externally operated signals to be connected with endogenous regulation circuits such as those that govern stress responses or phase changes, etc. / Diego Orzáez
The examples cited above are not limited to transferring a single transgene or a small modification into the genetic code of plants, rather they provide genetic circuits and complex metabolic pathways involving many genes. Some authors argue that, in order to achieve the levels of sophistication required to design such plants, engineering principles similar to those that allowed the industrial and information engineering revolutions must also be incorporated into biotechnology, including standardisation, modularity, and abstraction of function. These aspects are addressed by the discipline known as synthetic biology, which is opening up new horizons in plant biotechnology. The example of plants producing sexual pheromones was initially suggested and has already been partially implemented by a team of students at the Polytechnic University of Valencia in collaboration with the Spanish National Science Research Council (CSIC in its Spanish abbreviation), within the context of the iGEM synthetic biology project started in Boston in 2015. This work has also seeded other similar research projects, namely the sustainable bioproduction of insect pheromones in plants as an alternative to using synthetic pesticides.
«Today we know many of the genetic factors that allowed the domestication of our crop species»
On a different note, the example of the response to weather forecasts illustrates the potential of designing genetic circuits that would allow scientists to externally operate endogenous processes of agronomic interest, such as the flowering time, activation of defence mechanisms, or protection against environmental stress (Figure 5). Similar to the way that augmented reality or «cyborg» interfacing is starting to allow humans to surpass our innate capabilities within the outside environment, synthetic biology seeks to create crop plants with augmented capabilities that can produce more with less input, and in a more sustainable and environmentally friendly way.
The challenges are enormous but, for knowledge ecosystems in which the promotion of creativity is combined with the incentive of necessity, almost no technological challenge is insurmountable. The plants of the future will probably end up being the way we want them to be. Decisions are already starting to be made In this respect, and our present choices will shape the future of the next generations on Earth.
References
Barton, K. A., Binns, A. N., Matzke, A. J., & Chilton, M. D. (1983). Regeneration of intact tobacco plants containing full length copies of genetically engineered T-DNA, and transmission of T-DNA to R1 progeny. Cell, 32(4), 1033–1043. http://doi.org/10.1016/0092-8674(83)90288-x
Engler, C., Kandzia, R., & Marillonnet, S. (2008). A one pot, one step, precision cloning method with high throughput capability. PLOS ONE, 3(11), e3647. http://doi.org/10.1371/journal.pone.0003647
FAO. (2018). The future of food and agriculture: Alternative pathways to 2050. Food and Agriculture Organitzation of the United Nations. http://www.fao.org/3/I8429EN/i8429en.pdf
Fuentes, I., Stegemann, S., Golczyk, H., Karcher, D., & Bock, R. (2014). Horizontal genome transfer as an asexual path to the formation of new species. Nature, 511(7508), 232–235. http://doi.org/10.1038/nature13291
Herrera-Estrella, L., Depicker, A., Van Montagu, M., & Schell, J. (1983). Expression of chimaeric genes transferred into plant cells using a Ti-plasmid-derived vector. Nature, 303, 209–213. http://doi.org/10.1038/303209a0
Khan, M. Z., Zaidi, S. S., Amin, I., & Mansoor, S. (2019). A CRISPR way for fast-forward crop domestication. Trends in Plant Science, 24(4), 293–296. http://doi.org/10.1016/j.tplants.2019.01.011
Lander, E. S. (2016). The heroes of CRISPR. Cell, 164(1-2), 18–28. http://doi.org/10.1016/j.cell.2015.12.041
Mojica, F. J., Díez-Villaseñor, C., García-Martínez, J., & Almendros, C. (2009). Short motif sequences determine the targets of the prokaryotic CRISPR defence system. Microbiology, 155(3), 733–740. http://doi.org/10.1099/mic.0.023960-0
Pillet, S., Aubin, É., Trépanier, S., Bussière, D., Dargis, M., Poulin, J.-F., Yassine-Diab, B., Ward, B. J., & Landry, N. (2016). A plant-derived quadrivalent virus like particle influenza vaccine induces cross-reactive antibody and T cell response in healthy adults. Clinical Immunology, 168, 72–87. http://doi.org/10.1016/j.clim.2016.03.008
Qiu, X., Wong, G., Audet, J., Bello, A., Fernando, L., Alimonti, J. B., Fausther-Bovendo, H., Wei, H., Aviles, J., Hiatt, E., Johnson, A., Morton, J., Swope, K., Bohorov, O., Bohorova, N., Goodman, C., Kim, D., Pauly, M. H., Velasco, J., … Kobinger, G. P. (2014). Reversion of advanced Ebola virus disease in nonhuman primates with ZMapp. Nature, 514(7520), 47–53. http://doi.org/10.1038/nature13777
Schmutz, J., Cannon, S. B., Schlueter, J., Ma, J., Mitros, T., Nelson, W., Hyten, D. L., Song, Q., Thelen, J. J., Cheng, J., Xu, D., Hellsten, U., May, G. D., Yu, Y., Sakurai, T., Umezawa, T., Bhattacharyya, M. K., Sandhu, D., Valliyodan, B., … Jackson, S. A. (2010). Genome sequence of the palaeopolyploid soybean. Nature, 463, 178–183. http://doi.org/10.1038/nature08670
Stephan, A., Hahn-Löbmann, S., Rosche, F., Buchholz, M., Giritch, A., & Gleba, Y. (2017). Simple purification of Nicotiana benthamiana-produced recombinant colicins: High-yield recovery of purified proteins with minimum alkaloid content supports the suitability of the host for manufacturing food additives. International Journal of Molecular Sciences, 19(1), 95. http://doi.org/10.3390/ijms19010095
Vazquez-Vilar, M., Orzaez, D., & Patron, N. (2018). DNA assembly standards: Setting the low-level programming code for plant biotechnology. Plant Science, 273, 33–41. http://doi.org/10.1016/j.plantsci.2018.02.024
Wang, L., Jiang, S., Chen, C., He, W., Wu, X., Wang, F., Tong, T., Zou, X., Li, Z., Luo, J., Deng, Z., & Chen, S. (2018). Synthetic genomics: From DNA synthesis to genome design. Angewandte Chemie International Edition English, 57(7), 1748–1756. http://doi.org/10.1002/anie.201708741
Zsögön, A., Cermák, T., Naves, E. R., Notini, M. M., Edel, K. H., Weinl, S., Freschi, L., Voytas, D. F., Kudla, J., & Peres, L. E. P. (2018). De novo domestication of wild tomato using genome editing. Nature Biotechnology, 36, 1211–1216. http://doi.org/10.1038/nbt.4272