Vaccines and evolution
Why is it important to understand the genetic diversity of pathogens?
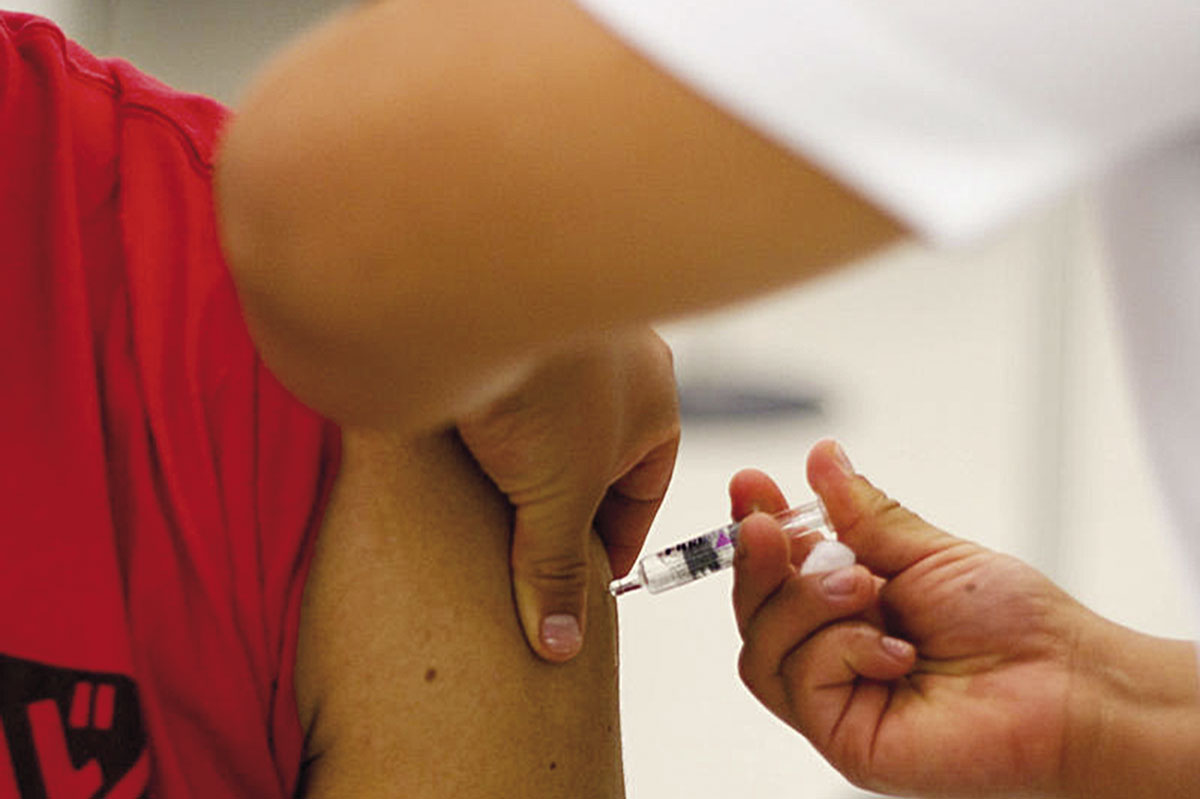
From the perspective of public health intervention, there is no better weapon than one which can prevent disease transmission or onset. Within the field of infectious diseases, vaccines have become the weapon that can control many of them. Today, however, there are a large number of emerging infectious diseases for which no vaccines exist, or long forgotten diseases that are re-emerging. New vaccines are being developed to attack the pathogens associated with them. However, despite the importance of good vaccine design, the genetic diversity of pathogens is not always taken into account. Studying their diversity helps identify how pathogens adapt to pressure from the immune system and to what extent the immune response benefits the host or, conversely, benefits the pathogen. Understanding these relationships will help us predict how effective and universal a vaccine can be, and thus aid in their design.
Keywords: antigenic variation, tuberculosis, flu, immune system, infectious diseases.
A brief history of vaccines
Infectious diseases represent a major scourge of mankind. Living in the 21st century makes it difficult to realize the economic, political and military impact they have had on our history. In addition to the well-known cases of smallpox epidemics in the New World, which wiped out 25% of the Aztec population in less than one year, or the death of twenty-four million Europeans during the years of the Black Death, throughout history many other outbreaks have taken their toll. The Panama Canal project was initially undertaken by France, who had to abandon it partially due to the high incidence of yellow fever and malaria among workers. In fact, during its construction by the United States of America, 85% of Panama Canal workers were hospitalized at some point due to these diseases. The construction of the canal was only made possible thanks to preventive measures taken against the respective vectors, which had just been identified in Cuba in the years after the war with Spain. Another case was the so-called Spanish flu, which killed 3% of the world population between 1918 and 1920, five times more deaths than those caused by the Great War, which had just ended. Many historians blame different plagues on the deterioration of many cities in the Roman Empire, contributing to its fall. In fact the so-called Antonine plague (165-185 AD), the causal agent of which is unknown (it is speculated that it was smallpox), is believed to have killed five million inhabitants of the Roman Empire. In a similar event, the Justinian plague struck the Byzantine Empire (541-542 AD). Napoleon himself had all his army vaccinated against smallpox, and publicly praised Edward Jenner for the development of the vaccine, despite being a citizen of an enemy country.
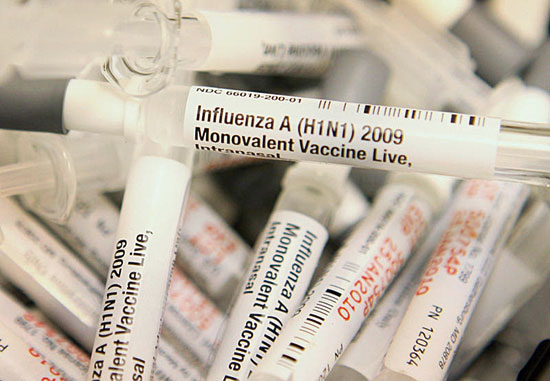
The seasonal influenza virus is of type H3N2. But this is only since 1968, when the H3N2 subtype displaced the previous one (H2N2). Additionally, many other less frequent types still circulate today (H1N1, H1N2). / Agència SINC
The causes of many infectious diseases only began to be understood when, in 1680, the Royal Society of London publicly acknowledged the observations of the Dutch scientist Antonie van Leeuwenhoek: the existence of single-celled microscopic organisms. That «entities» could be the cause of major epidemics had been proposed even before Leeuwenhoek’s observations, by the likes of Girolamo Fracastoro. But it was the work of the Dutch scientist, and later of academics such as Semmelweis (in the field of hygiene) and John Snow (in the field of epidemiology), which served to support the formulation of the so-called «germ theory». This theory postulated that many of the most common diseases were actually due to infections by microorganisms, and gave definitive support to the work of Louis Pasteur and Robert Koch. Many of the causal agents of the largest epidemics were described in the nineteenth or early twentieth century, and their identification led to an era of innovation in the prevention, diagnosis and treatment of infectious diseases (De Kruif, 1926). As a result of this, the second half of the twentieth century was characterized, if not by the eradication, at least by the displacement of infectious diseases as the leading cause of morbidity in developed countries. In this sense there is a general consensus that the development of vaccines and mass vaccination campaigns, many run worldwide, have saved millions of lives, and virtually eradicated many diseases (Nabel, 2013). Throughout the history of vaccine research, a central role has been occupied by a disease that is responsible for the deaths of hundreds of millions of people: smallpox.
Contrary to popular belief, the process of inoculating pathogens into people to generate a protective immune response to future exposures did not begin with Edward Jenner, the British physician to whom the first vaccine is justly attributed. The first cases of smallpox inoculation are attributed to a Buddhist monk around 1020 AD. It was recorded that he sampled wound remains from smallpox patients and inserted them in the nostrils of people without the disease, thus granting a degree of protection against new infections. In eighteenth century England, the bovine version of the disease was as important economically as the human version was deadly. Working with both types, Edward Jenner successfully developed and tested the first vaccine against smallpox in history, using the bovine version of the disease. The vaccine was not only a scientific breakthrough that led the way for many others of its kind, but also marked the beginning of public health activities on a large scale, with smallpox being the greatest example of disease eradication resulting from coordinated mass vaccination campaigns. In 1977, Ali Maow Maalin, aged 23, a resident of Merca (Somalia), became the last official case of smallpox in the world and the first great success of the World Health Organization (WHO) in the eradication of infectious diseases. Success with smallpox vaccination and varying degrees of success with diseases such as polio, measles, mumps or diphtheria have not prevented, however, the fact that even today many infectious diseases remain common, or have resurfaced, creating the need for a new approach to developing vaccine.
«Many of the causal agents responsible for the largest outbreaks were described in the nineteenth or early twentieth century. Their identification led to an era of innovation in prevention, diagnosis and treatment»
The pathogen’s perspective
A possible cause of failure in vaccine design is that, in many cases, they do not take into account the genetic diversity of the pathogen or its ability to adapt to new selection pressures, such as vaccinations or antibiotics. From the genetic standpoint, pathogens, like human populations, are not homogeneous, but have accumulated many changes throughout their evolutionary history. There are pathogens with high genetic diversity, and some even have pathogenic and non-pathogenic strains, like Escherichia coli. There are also highly specialized pathogens with a very low range, as in the case of leprosy and tuberculosis bacilli or plague-related bacteria. Some have high recombination rates, such as Neisseria meningitidis and some viruses, whereas others are genetically monomorphic. In the context of vaccines, this existing variation must be taken into account, as well as the ability of the pathogen to mutate and find variations allowing them to escape from the action of the immune system, or from the protection conferred by vaccines. All these aspects can affect pathogen evolution, new vaccine development and universality. To illustrate the impact of developments in vaccine design, let us look at two very different examples: the case of the flu vaccine, and the vaccine for tuberculosis.
Success with smallpox vaccination and other diseases has not prevented, however, the fact that even today many infectious diseases remain common, or have resurfaced, creating the need for a new approach to developing vaccine. / Aki Hänninen
Influenza: evading the immune response
Viruses in general, and RNA-based viruses in particular, are typical examples of organisms evolving so fast that they are able to find solutions to many of the challenges we put in their path during prevention or treatment. Their high rates of mutation and/or recombination are not the only secret to their success, but also their large population sizes, allowing them to find the most optimal evolutionary solutions faster. This high variability has direct consequences on the design of drugs and vaccines to treat or prevent infection. A paradigmatic case, perhaps the most studied, is the vaccine used each year for seasonal flu.
Birds are the natural reservoir of all influenza viruses, whether seasonal or epidemic. Influenza viruses that affect humans are known as type A and are generally differentiated based on the combination of two proteins, HA (haemagglutinin) and NA (neuraminidase). These proteins are recognized by the immune system, in other words, they are antigens. Every major recombination event that has generated new combinations of these two proteins has been accompanied by a radical change in their antigenicity, resulting in pandemic episodes of greater or lesser relevance. This is what happened with the 1918 virus, which killed an estimated 50 million people globally, commonly known as the Spanish flu virus. For example, the influenza virus that affects us every year (also known as seasonal influenza) is of type H3N2. But this is only since 1968, when the H3N2 subtype displaced the previous one (H2N2). Additionally, many other less frequent types still circulate today (H1N1, H1N2).
«The smallpox vaccine was not only a scientific breakthrough paving the way for many others of its kind, but also marked the beginning of public health research on a large scale»
From the above, it is clear that the influenza virus is highly skilled at generating antigenic diversity. In fact, attending to the mechanism involved and especially to the consequences of the accumulation of such variation, one can differentiate between episodes of «antigenic drift» and episodes of «antigenic shift» (Rambaut et al., 2008). Previously I have referred to the latter, usually connected to a radical change in the antigenic composition of the virus. Because the general population has not been exposed previously to this new combination, the result is the outbreak of pandemics of varying severity. Historically we distinguish between five major pandemics, including the most recent, known as swine flu, caused by H1N1 type virus.
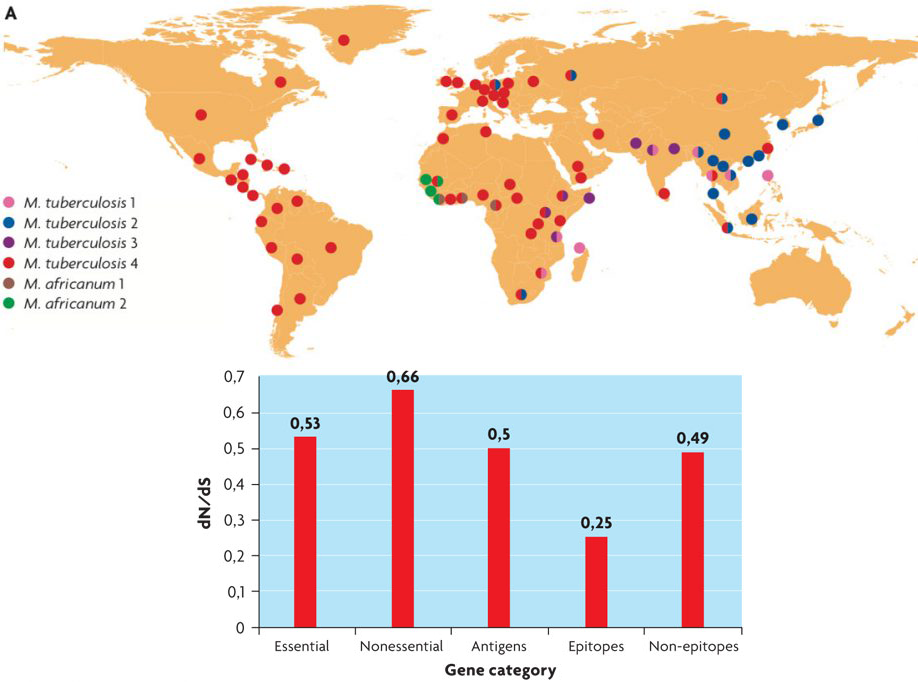
A) Global distribution of major lineages within the tuberculosis complex. The colours indicate the dominant lineage in the different geographical areas. / From Comas & Gagneux, 2009. B) Known epitopes of M. tuberculosis are the most conserved part of the pathogen’s genome, suggesting that tuberculosis does not use a strategy of immune evasion but takes advantage of it for transmission / Adapted from Comas et al., 2010
Contrary to what happens when the virus completely changes its antigenic composition, seasonal flu must face the fact that infection of a host results in some degree of protective immunity, which will limit the number of people susceptible to the same type of virus each year. The way in which the virus evades populations with acquired immunity is by bringing together its high mutation rate and population size, allowing it to find amino acid changes that circumvent immunity acquired from previous infections. The result of this is that the seasonal influenza virus is slightly different each year due to the accumulation of changes made to escape the immune system (Figure 1). This constant change is what causes the recommended vaccine against influenza to be different every year. An expert committee annually decides on the best combination of antigens to generate general immunity, based on data for the types of influenza circulating in previous years. For instance, in 2013 the WHO has recommended that the vaccine be composed of a mixture of antigens from a strain of H1N1 detected in 2009, another strain H3N2 from 2011, and one for influenza type B from 2010, which is less common, but present in many countries.
«Seasonal influenza virus differs slightly each year due to accumulated changes for escaping the immune system. This constant change means the vaccine against influenza must also change every year»
Tuberculosis: manipulating the immune response
The tuberculosis bacillus constitutes a special case in terms of the impact of pathogen diversity on the immune response. Between 1908 and 1921 two microbiologists at the Pasteur Institute, Calmette and Guérin, isolated and serially cultured the bovine tuberculosis bacillus, to create an attenuated strain that could be used to immunize against tuberculosis infection in humans. This attenuated strain is known as the Bacillus Calmette-Guérin (BCG) and is the most widely used vaccine in history. The bovine version of tuberculosis is caused by Mycobacterium bovis which is part of what is known as the tuberculosis complex, together with other bacilli which cause tuberculosis in animals, but rarely in humans, and the bacilli that practically only cause human tuberculosis (Mycobacterium tuberculosis and Mycobacterium africanum). Although they all have different species names, their genome-level differences are so small (about 2,000 nucleotide changes maximum) that today they would be considered a single species. When Calmette and Guérin developed the vaccine from a virulent strain of bovine tuberculosis, they were unaware that during the passaging process in the laboratory (on sliced potato supplemented with glycerol) a series of changes at the genomic level took place, which were increased when the vaccine strain was distributed to laboratories around the world. Because at that time there were no methods to keep samples frozen, to maintain them, laboratories performed cell passaging or sub-cultures. The result is that different vaccine strains accumulated different mutations in their genome, resulting in a pedigree that enables the evolution of the vaccine to be traced over the past ninety years. Thus, the tuberculosis vaccine inadvertently became an experiment in evolution that lasted decades. However, the tuberculosis vaccine is not effective against the most common and contagious disease, which is pulmonary tuberculosis in adults. However, it is effective against extra-pulmonary forms of the disease in children, and hence the mass vaccination campaigns carried out with BCG in countries with high incidence of this kind of tuberculosis.
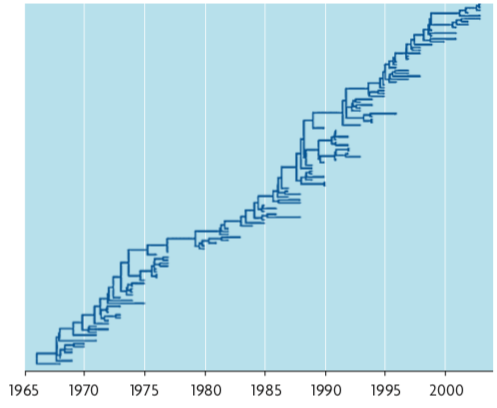
Figure 1. Phylogenetic tree of a region of the HA gene of influenza A (H3N2) from virus samples obtained between 1968 and 2002. The tree reflects a predominant lineage from which secondary branches derive that last a few years. The central lineage reflects the adaptation of the virus antigen over time in response to the immune system. / Source: PLoS Comput Biol
Finding an effective vaccine against pulmonary tuberculosis has become one of the main objectives of the WHO in the last 30 years. Although TB can be cured if treatment is followed, it is estimated that one third of the world’s population is latently infected and about 10% have symptoms of active TB at some point in their lives. This indicates that although the active cases can be treated with antibiotics, in terms of cost-effectiveness it would be much more worthwhile to implement a vaccine that could prevent infection or at least impede the re-activation of latent infections. More than 10 vaccine candidates are currently in various stages of development, some of them being tested in human clinical trials. However, TB is a complex disease of whose causative agent, despite having been discovered over one hundred years ago, we still know little about, as we do of the interaction between the bacteria and the immune system during infection. The advent of genomic sequencing, and more recently of massive sequencing technologies, is enabling an understanding of the full extent this pathogen’s diversity, which has led to new insights into transmission, evolution and, regarding the topic at hand, immune response.
«The tuberculosis vaccine inadvertently became an experiment in evolution that lasted decades. This vaccine, however, is not effective against the most common and contagious variation of tuberculosis bacillus»
As seen in the case of influenza, most pathogens, in one way or another, accumulate changes in proteins or areas of the genome which are recognized by the immune system during infection. Such proteins are known as antigens and their accumulated changes are associated to antigenic variation which allows the pathogen to evade human defences. However, the case of tuberculosis is special. A study using massive sequencing technologies on a set of strains representing the known diversity of the bacillus has revealed that the antigens, and specifically the peptide fragments recognized by the immune system known as epitopes, are hyper-conserved in Mycobacterium tuberculosis (Comas et al., 2010). In fact when compared with essential genes, pathogen genes that cannot be eliminated because they are necessary for survival, the epitopes are equally or better preserved and accumulate fewer amino-acid changes than non-essential genes or other areas not recognized by the immune system. However, from experiments in mice we know that the adaptive immune response, and especially that mediated by T CD4+ cells, is essential to control the infection. Why does tuberculosis bacillus not try to evade this response by taking advantage of changes in the sequence of its epitopes? The simplest answer is because, in the case of tuberculosis, the pathogen needs the immune response to be transmitted. The immune response allows the bacillus to complete its cycle of infection, first by inducing a state of latency and then overreacting in the 5-10% of cases that the bacterium is reactivated. The inflammatory reaction is in fact part of what causes damage to lung tissues, which are expelled along with the pathogen during sneezing, thus allowing transmission to occur. This life cycle indicates that although the host benefits from the immune response, this reaction also appears to benefit the pathogen. In fact, HIV-positive patients with low CD4+ levels, contract extra-pulmonary forms more frequently, non-transmissible disease forms, with lower bacillary load in the sputum. This indicates that a proper immune response is needed to contain the infection in the lung in the first place, and for transmission in cases where the bacteria are re-activated from a latent infection.
The tuberculosis vaccine is not effective against the most common and contagious disease, which is pulmonary tuberculosis in adults. However, it is effective against extra-pulmonary forms of the disease in children, and hence the mass vaccination campaigns carried out with BCG in countries with high incidence of this kind of tuberculosis. Above, an immunisation campaign in Delhi (India). / OMS/P. Virot
These results shed light on why BCG vaccine efficacy is so irregular, since the bovine form of the bacillus is not essentially very different from the human tubercle bacillus at the genomic level. In fact, it is a red flag to vaccine designers, warning them about the use of some hyper-conserved epitopes, because they may partially favour the pathogen, not the host. Therefore, in contrast to other approaches to vaccine design, which aim to identify the most conserved regions of the genome (known as reverse vaccinology), for tuberculosis and other infections, maybe new antigens should be sought. These should be based on hypervariable regions of the genome with an accumulation of variations that help to escape from the immune response. Strategies similar to that employed by Mycobacterium tuberculosis, using part of the immune response to complete its life cycle, have recently been described for other pathogens such as HIV or Streptococcus pneumoniae.
The two cases analysed here, as well as many others, lead to a key consideration. Variation exists in a pathogen and several questions must be considered in the design of new vaccines: What is the nature of pre-existing variation? Is this variation linked to immunity-related or vaccination-related selective pressures? What is the pathogen’s ability to adapt to new selection pressures? Answering these questions will lead to better vaccine design, through a richer understanding of the interaction between the immune system and the pathogen, enabling us to distinguish between those elements that are part of the protective immune response from those interactions that can benefit the pathogen.
REFERENCES
Comas, I. and S. Gagneux, 2009. «The Past and Future of Tuberculosis Research». PLoS Pathogens, 5(10): e1000600. DOI: <10.1371/journal.ppat.1000600>.
Comas, I. et al., 2010. «Human T Cell Epitopes of Mycobacterium tuberculosis Are Evolutionarily Hyperconserved». Nature Genetics, 42(6): 498-503. DOI: <10.1038/ng.590>.
De Kruif, P., 1926. Microbe Hunters. Harcourt, Brace and Co. Nova York.
Nabel, G. J., 2013. «Designing Tomorrow’s Vaccines». The New England Journal of Medicine, 368(6): 551-560.
DOI: <10.1056/NEJMra1204186>.
Rambaut, A. et al., 2008. «The Genomic and Epidemiological Dynamics of Human Influenza A Virus». Nature, 453(7195): 615-619. DOI: <10.1038/nature06945>.