On the Trail of HIV
Evolutionary studies and understanding of AIDS

|
ABSTRACT Keywords: phylogeny, biodiversity, genetic variation, mutation, public health, virus. What is HIV and what is its significance? Acquired immune deficiency syndrome (AIDS) is a global public health problem. Since its discovery in 1983, the human immunodeficiency virus (HIV) has killed about 30 million people and more than 40 million people are infected, most in developing countries. The human immunodeficiency virus is a retrovirus that specifically infects immune system cells expressing the membrane receptor CD4, mainly T4 lymphocytes. As a result, the virus causes gradual weakening of the immune system. In advanced stages, the infection causes the appearance of opportunistic diseases and cancers characteristic of AIDS. Without treatment, death occurs on average about 10 years after infection. However, there is great variability in the progression time of the disease, depending on the characteristics of the virus and the patient’s immune system. The study of the evolution of HIV has shed light not only on its natural history but also on its pathogenesis mechanisms. Through evolutionary studies we can attempt to predict, for example, how the virus might respond to the presence of drugs or which variants are more likely to spread in the future. In fact, evolutionary studies have long had a positive impact on the design of treatments for human disease in general. For example, estimates for the mutation rate (probability of changes in genetic material) of HIV and hepatitis C have shown that all possible changes in a single nucleotide occur daily inside a patient. Therefore, drug resistance is almost inevitable unless a combination of therapies is used, each with different targets, and thus requiring multiple genetic changes in the virus for it to develop resistance to the treatment.
|
«Evolutionary studies can help predict how the virus might respond to drugs or which variants are more likely to spread in the future» | |
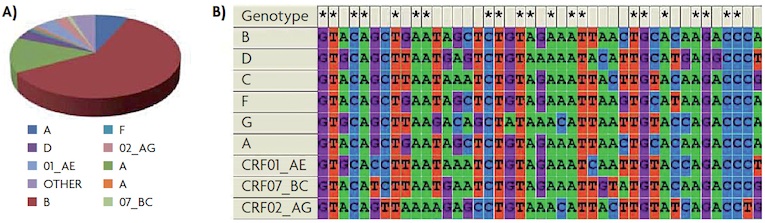 Marianoel Pereira-Gómez, Joan B. Peris i Rafael Sanjuán Marianoel Pereira-Gómez, Joan B. Peris i Rafael SanjuánFigure 1. A) Abundance of HIV-1 different viral genotypes in the world. HIV-1 is classified into groups (M, N, O and P), group M being the most numerous; within this group are the subtypes A, B, C, D, F and G. Several other recombinant forms have been described circulating in human populations, the most abundant worldwide are 01_AE, 02_AG and 07_BC. B) Sequence alignment of different subtypes of HIV-1 in a region from gene that codifies for the envelope. Asterisks mark the conserved sites. The great divergence can be easily observed. |
||
|
Origins and epidemiology Studies on variability and evolution of HIV have shown us that there are two types, HIV-1 and HIV-2, with different geographical origin and related to different simian immunodeficiency viruses (SIV). Both originated as the result of zoonosis, that is, transmission of a disease from animals to humans. HIV-1 originated in chimpanzees after one or more recombination events between different SIV, allowing the disease to jump the species barrier due to environmental and/or genetic factors (Gao et al., 1999). On the other hand, HIV-2 probably originates from the VIS of the grey mangabe. HIV-1 causes most infections and is the more virulent of the two types. The divergence observed in the whole genome between HIV-1 and HIV-2 is approximately of 40% to 60%. Between different subtypes of HIV-1, divergence can be up to 35%, while in the so-called M group (which causes 95% of HIV infections) it is estimated between 10% and 30%. The highest divergence values have been observed in the gene which codes for the envelope (env) (Brenner, 2007). Figure 1 shows the abundance of the different genotypes of HIV-1 in the world and an example of diversity in the env gene between the different genotypes of HIV-1. Epidemiology is devoted to studying the causes, incidence, distribution and control of diseases in human populations. From a practical point of view it is important to consider both the epidemic process and the evolution of pathogens, whether related to the evolution of drug resistance and/or virulence of the pathogen, vaccine design, or the emergence of new pathogens and diseases. Understanding how a pathogen can successfully invade and spread in the population is of paramount interest. The 2010 UNAIDS report showed that new HIV infections have been declining since the late 90s in most of the affected countries in sub-Saharan Africa, due to preventive measures like the use of condoms. Conversely, in some countries in Eastern Europe and Central Asia, HIV incidence has increased. The report also showed how increased access to treatment has resulted in a significant decrease in deaths from HIV, and how vertical transmission from mother to child can be virtually eliminated if treatment is applied in the first weeks of pregnancy. |
«Transmission of the virus from mother to child can be virtually eliminated if treatment is applied in the first weeks of pregnancy» | |
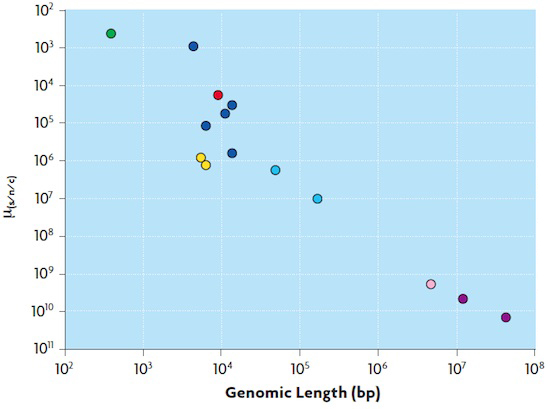 Marianoel Pereira-Gómez, Joan B. Peris i Rafael SanjuánFigure 2. Relationship between mutation rate and genome size in different organisms. The mutation rate is expressed as nucleotide substitutions per generation (μ(s/n/c)) and genome size in bases or base pairs (bp). Green represents the estimate for a viroid, an infectious agent composed of a non-coding small RNA molecule. Blue represents RNA viruses; yellow, single stranded DNA viruses, and light blue the double-strand DNA virus. Grey shows the estimate for a model E. coli bacterium, and finally two estimates for eukaryotic organisms are represented. The estimated value for HIV-1 is displayed in red. Marianoel Pereira-Gómez, Joan B. Peris i Rafael SanjuánFigure 2. Relationship between mutation rate and genome size in different organisms. The mutation rate is expressed as nucleotide substitutions per generation (μ(s/n/c)) and genome size in bases or base pairs (bp). Green represents the estimate for a viroid, an infectious agent composed of a non-coding small RNA molecule. Blue represents RNA viruses; yellow, single stranded DNA viruses, and light blue the double-strand DNA virus. Grey shows the estimate for a model E. coli bacterium, and finally two estimates for eukaryotic organisms are represented. The estimated value for HIV-1 is displayed in red. |
||
|
Evolutionary mechanisms Mutation creates new genetic variability, while recombination, broadly defined, produces high genetic variability. Deterministic factors (such as mutation, migration or selection) or random factors (such as genetic drift) can act on this genetic variability. Some viruses use RNA as genetic material instead of DNA, and the former are characterized by particularly high mutation rates. This is the case of HIV, which uses a special polymerase to replicate its genome. This polymerase is called reverse transcriptase, and it copies RNA to make DNA. Most polymerases that use RNA as a template lack error correction mechanisms and this leads to mutations, which favour viral populations with high genetic variability. Figure 2 shows how mutation rates vary in different organisms. The highest mutation rate was described in a plant viroid. Within the virus group, RNA viruses have the highest mutation rates, followed by single strand DNA (ssDNA) viruses and double stranded DNA viruses. The mutation rate of HIV is within the range typical for RNA viruses. Organisms with a cellular structure have mutation rates about 100,000 times lower than those of RNA viruses. Another very important aspect in the evolution of HIV is recombination. The HIV virion possesses two genome copies as RNA and the reverse transcriptase can jump from one strand to another during DNA synthesis, resulting in recombinant molecules (Simon-Lorière and Holmes, 2011). Recombination played a key role in the origins of HIV and could result in a new type of HIV in the future. Recombination affects host range expansion, virus virulence, immune system evasion or acquisition of drug resistance. For example, the circulating recombinant form 01_AE, once settled in northern Thailand, increased its frequency and quickly dominated the rest of Thailand, Laos, Cambodia and Vietnam (Stephens, 2005). Genetic drift is the process by which allele frequencies change randomly from one generation to the next due to sampling effects in populations. The smaller the population size the greater its intensity, and in the case of HIV, it occurs mainly in the transmission between people, because it entails bottlenecks in the viral population size. There is consensus that HIV infection usually begins with the transmission of one or a few infectious particles (Keele et al., 2008). Bottlenecks may also occur in the patient, for example in tissue subdivisions. On the other hand, natural selection causes changes in allele frequencies by acting on mutations and/or genomes that influence biological efficiency. Selection may be adaptive or positive when it promotes the fixation of mutations that are beneficial in a given environment or negative when deleterious mutations are eliminated. Both the host’s immune system and antiviral treatments exert strong selection pressures on the virus, as discussed below. Evolutionary mechanisms acting on viral populations may have a different role in the host and between hosts. For example, when observing the phylogenetic relationships of viral variants for the env gene encoding the viral envelope, one can distinguish two different patterns. On the one hand, intra-host phylogenies indicate the importance of the action of selection pressures while, on the other, inter-host phylogenetic trees suggest that genetic drift plays a greater role (Grenfell et al., 2004) (Figure 3). |
«Recombination played a very important part in the origin of HIV and could lead to a new type of HIV in the future» | |
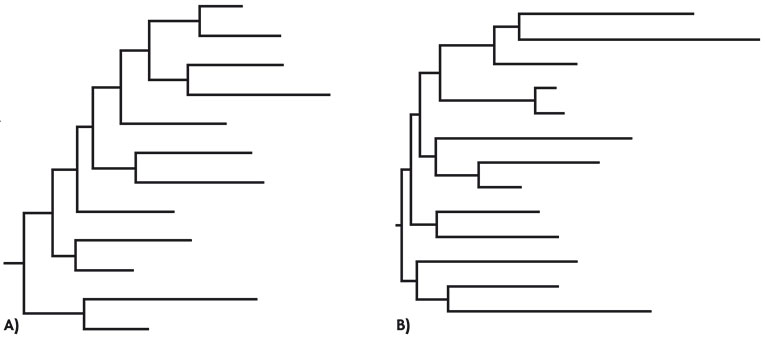 Marianoel Pereira-Gómez, Joan B. Peris i Rafael Sanjuán Marianoel Pereira-Gómez, Joan B. Peris i Rafael SanjuánFigure 3. Dynamics in the evolution of HIV env gene. A) Intra-host phylogeny: the structure of the tree’s branches is similar to a ladder and indicates the continuous action of a selection process produced by the action of the immune system or the presence of antivirals, in fixing those mutations that are favourable and making extinct those lineages without them. B) Inter-host phylogeny: in this case the image shows the persistence of various lineages over time. The shape of the tree is mainly determined by neutral demographic processes. |
||
|
Escaping the immune system The host’s immune system has an innate component (non-specific) and another adaptive component (specific). The innate immune response is characterized by fever, inflammation and other symptoms, which are controlled by processes such as cytokine secretion (e.g., interferon). In the case of HIV, the enzyme APOBEC3G (a protein that edits single-stranded DNA) is involved in the innate immune response and causes mutations that inactivate the virus during the process of viral DNA synthesis from RNA. To counteract the effect, HIV incorporated a gene into its genome called viral infectivity factor (vif) (Kirchhoff, 2010). In the case of specific or adaptive immune response in vertebrates, there are a number of genes coding for the major histocompatibility complex (MHC). This complex presents portions of viral proteins called epitopes, allowing the launch of an immune response against the virus. Human populations are extraordinarily variable for MHC genes and therefore recognize different viral epitopes. Some MHC variants have been shown to respond better to infection than others, and thus are associated with a slower progression of the disease (Buckheit III et al., 2012). In order to understand this, it becomes necessary to consider the evolution of HIV within patients. The pressure exerted by specific immune responses selects for viral mutants able to evade this response. However, these mutations may compromise the ability of the virus to carry out its infectious cycle, if the mutations happen to affect important regions of the viral proteins (conserved regions). Precisely those MHC variants capable of recognizing viral epitopes in conserved regions are the ones associated with a better prognosis. |
||
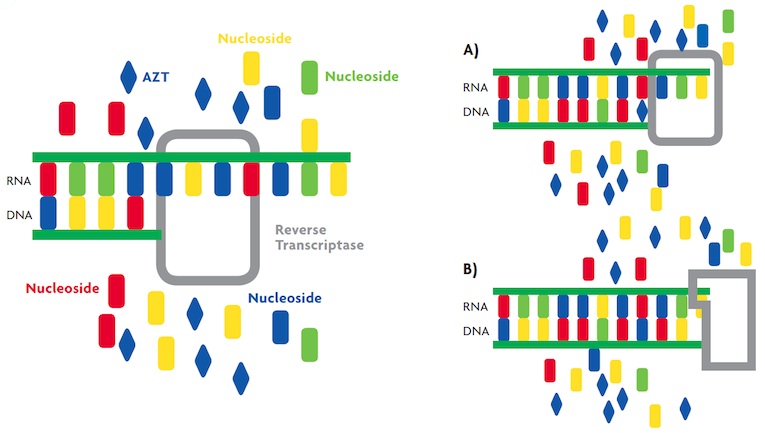 Marianoel Pereira-Gómez, Joan B. Peris i Rafael Sanjuán Marianoel Pereira-Gómez, Joan B. Peris i Rafael SanjuánFigure 4. Reverse transcriptase action mechanism. A) Figure representing the incorporation of AZT (blue diamond), a nucleoside analogue, in viral DNA synthesis by the reverse transcriptase, which interrupts synthesis. B) When a resistance mutation occurs in the active site of the reverse transcriptase (for example, T215Y), the mutant enzyme discriminates AZT and can continue viral DNA synthesis. |
||
|
Antiviral resistance Because viruses are obligate intracellular parasites and use the cellular machinery, it is difficult to find compounds that only affect viral activity, because they can also adversely affect the host cell. Therefore, an essential approach to the design of new drugs is to identify the specific steps of the viral infectious cycle. The reverse transcriptase and the protease responsible for processing the polyprotein encoded by HIV are ideal targets, as they are unique to the virus. The first compound that showed effectiveness against the reverse transcriptase was azidothymidine (AZT), a nucleoside analogue whose structure is similar to that of the nucleotides that make up DNA. When AZT is present in the cell, the reverse transcriptase erroneously adds it to the newly synthesized DNA strand and interrupts synthesis (Figure 4). However, it was observed that after one or two years of treatment, HIV patients became unresponsive and the disease continued its progression. The resistance was due to mutations in the reverse transcriptase gene, which were convergent, meaning that they appeared in viral populations from different patients. To date, some of the mutations described are located in the active site of reverse transcriptase, and are named L74V and T215Y (nomenclature indicates the amino acid change, and the position in which it occurs) (Bennett et al., 2009) (Table 1). Importantly, some mutations cause cross-resistance to other antiviral nucleoside analogues (Margeridon-Thermet and Shafer, 2010). Thus, the genetic composition of the HIV population changed over time in response to the presence of AZT. This is a clear example of evolution by natural selection. Consequently, the current treatment involves combinations of three or four drugs, including reverse transcriptase and protease inhibitors. This combination therapy is very effective at blocking virus replication and reducing viral load in the blood. However, as the virus is latent in some cells, the blockade is not complete, potentially allowing resistant viral particles to appear, preventing the eradication of the virus. |
«The pressure exerted by specific immune responses selects for viral mutants able to evade this response» | |
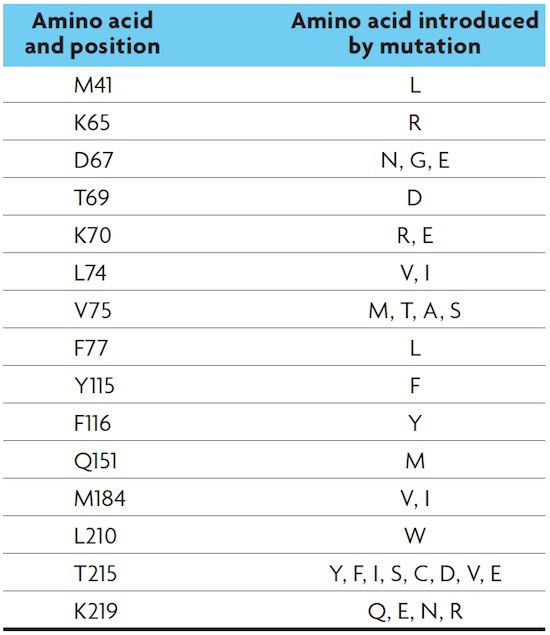 HIV Drug Resistance Database (<http://hivdb.stanford.edu/>) HIV Drug Resistance Database (<http://hivdb.stanford.edu/>)Table 1. Examples of resistance mutations to antiviral reverse-transcriptase-inhibitor nucleoside analogues. |
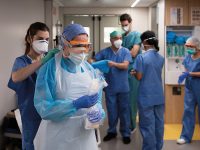
 Agencia SINC Agencia SINCHIV can cause the appearance of opportunistic diseases and cancers characteristic of AIDS. Without treatment, death occurs on average about 10 years after infection. However, there is great variability in the progression time of the disease. Above, human antibody of a patient infected with HIV. |
|
|
Vaccine design Vaccines induce a specific cellular and/or humoral immune response, so that the vaccinated individual develops an “immune memory” which gives a better chance in fighting off the pathogen. Traditionally different strategies have been used, such as inactivated pathogens (by heat or chemically), attenuated pathogens or administration of surface compounds that act as antigenic determinants that are recognized by the immune system’s cells. Vaccination has had great success in the prevention and control of various diseases such as polio, smallpox, measles, rubella, or flu. However, we have yet to see effective vaccines developed against HIV-1 or hepatitis C. One of the difficulties faced in developing a vaccine against HIV-1 is the high genetic variability and rapid evolution of the virus. As a consequence of this large variability, especially in the gene encoding the envelope protein, a vaccine that protects against a particular subtype may not grant, or grant only partial protection against other subtypes. However, epidemiological studies report that a low efficacy vaccine may reduce the incidence of HIV-1 by a significant percentage (UNAIDS, 2010). In this area, we highlight the study conducted in 2009 on the Thai population. The results of phase III clinical trials showed a 30% efficacy in protecting against HIV. The vaccine consisted of a combination of a vector expressing HIV-1 immunogens (substances that induce the adaptive immune response) and a recombinant version of the envelope protein (Rerks-Ngarm et al., 2009). Although this result does not provide sufficient evidence to back the general distribution of the vaccine, it gives clues about where further research should aim in order to enhance the immune response. New research should aim towards creating new constructions, use of multiple doses or even the administration of more and better adjuvants (immune response enhancers) that might be supplied along with it. |
«One of the difficulties faced when developing a vaccine against HIV-1 is the high genetic variability and rapid evolution of the virus» | |
 Jon Rawlinson Jon RawlinsonIncreased access to treatment has resulted in a significant decrease in deaths from HIV. Above, a sign in Zambia reminds the population about the importance of testing themselves for HIV. |
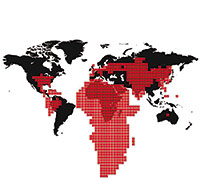 Agencia SINC Agencia SINCThe map, made in 2008, shows the number of infections (in red) per geographic area (in black). Each red square amounts to 50,000 infections. New HIV infections have been declining since the late 90s in most of the affected countries in sub-Saharan Africa, due to preventive measures like the use of condoms. Conversely, in some countries in Eastern Europe and Central Asia, HIV incidence has increased. |
|
|
Conclusions and future prospects Well aware of the importance of AIDS, many funds have been allocated to the study of HIV. The main objectives of the research groups engaged in these endeavours are to find effective antiviral drugs and a vaccine. The first objective has been largely fulfilled. However, obtaining an effective vaccine seems a goal with a distant horizon. The high genetic variability of HIV, due to its high mutation and recombination rates, along with the characteristic latent infection of the disease, makes it especially difficult to combat. Thus, knowledge and understanding of the evolutionary mechanisms that govern the evolution of HIV are key not only to the secrets of how the disease originated, but also how it develops, how to treat it, how it might change in the future and, ideally, how to eradicate it. |
«The objective of finding effective antiviral drugs has been largely fulfilled. However, obtaining an effective vaccine seems a goal with a distant horizon» | |
REFERENCES
Bennett, D. E. et al., 2009. «Drug Resistance Mutations for Surveillance of Transmitted HIV-1 Drug-Resistance: 2009 Update». PLOS One, 4: e4724. DOI: <10.1371/journal.pone.0004724>.
Brenner, B. G., 2007. «Resistance and Viral Subtypes: How Important Are the Differences and Why Do They Occur?». Current Opinion in HIV and AIDS, 2(2): 94-102. DOI: <10.1097/COH.0b013e32801682e2>.
Buckheit III, R. W. et al., 2013. «The Implications of Viral Reservoirs on the Elite Control of HIV-1 Infection». Cellular and Molecular Life Sciences, 70(6): 1009-1019. DOI <10.1007/s00018-012-1101-7>.
Gao, F. et al., 1999. «Origin of HIV-1 in the Chimpanzee Pan troglodytes troglodytes». Nature, 397: 436-441. DOI: <10.1038/17130>.
Grenfell, B. T. et al., 2004. «Unifying the Epidemiological and Evolutionary Dynamics of Pathogens». Science, 303(5656): 327-332. DOI: <10.1126/science.1090727>.
Keele, B. F. et al., 2008. «Identification and Characterization of Transmitted Early Founder Virus Envelopes in Primary HIV-1 Infection». PNAS, 105(21): 7552-7557. DOI: <10.1073/pnas.0802203105>.
Kirchhoff, F., 2010. «Immune Evasion and Counteraction of Restriction Factors by HIV-1 and Other Primate Lentiviruses». Cell Host and Microbe, 8(1): 55-67. DOI: <10.1016/j.chom.2010.06.004>.
Margeridon-Thermet, S. and R. W. Shafer, 2010. «Comparison of the Mechanisms of Drug Resistance Among HIV, Hepatitis B, and Hepatitis C». Viruses, 2(12): 2696-2739. DOI: <10.3390/v2122696>
Rerks-Ngarm, S. et al., 2009. «Vaccination with ALVAC and AIDSVAX to Prevent HIV-1 Infection in Thailand». The New England Journal of Medicine, 361: 2209-2220. DOI: <10.1056/NEJMoa0908492>.
Simon-Loriere, E. and E. C. Holmes, 2011. «Why do RNA Viruses Recombine?». Nature Reviews Microbiology, 9: 617-626. DOI: <10.1038/nrmicro2614>.
Stephens, H. A. F., 2005. «HIV-1 Diversity Versus HLA Class I Polymorphism». Trends in Immunology, 26(1): 41-47. DOI: <10.1016/j.it.2004.11.001>.
UNAIDS, 2010. Global Report. UNAIDS Report on the Global AIDS Epidemic. Available at: <http://www.unaids.org/en/media/unaids/contentassets/documents/unaidspublication/2010/JC1958_GlobalReport2010_full_es.pdf>.