Viruses against bacteria
The use of phages to fight bacterial resistance to antibiotics
An introduction to phages
Micro-organisms and humankind have coexisted since the origin of our species, resulting in both good and bad interactions. Many micro-organisms are beneficial to humans, while others can cause disease. Among the best known micro-organisms are bacteria and viruses. Unlike bacteria, which are single-celled prokaryotic organisms, viruses are obligate intracellular parasites, meaning that they need a host to complete their life cycle. They are made up of nucleic acids, which form their genome, and proteins to coat it. They are inert in the extracellular environment and can recognise their hosts very specifically. Once recognised, they inject their genetic material (i.e., their genome) into the host and hijack its cellular machinery in order to replicate and produce viral progeny. In general, viruses are capable of infecting organisms from all three domains of life – bacteria, archaea and eukaryotes – and are actually highly specific: each virus is specialised in recognising a small range of hosts. For example, viruses that infect bacteria, called bacteriophages, recognise only certain bacteria, and will never recognise or infect eukaryotic cells. Among eukaryotic viruses, plant viruses are very different from animal viruses, and among animals, there will be viruses that infect only one type of cell type.
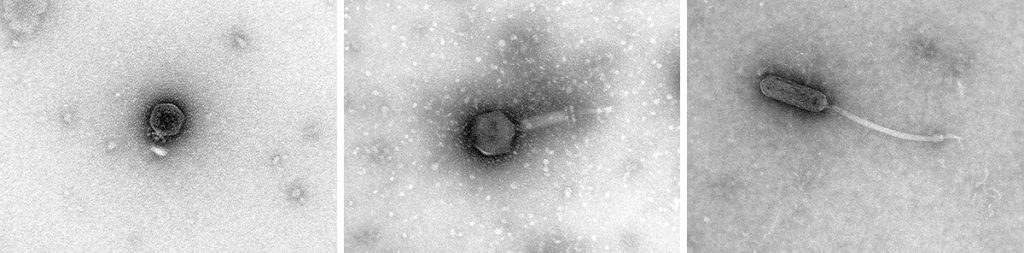
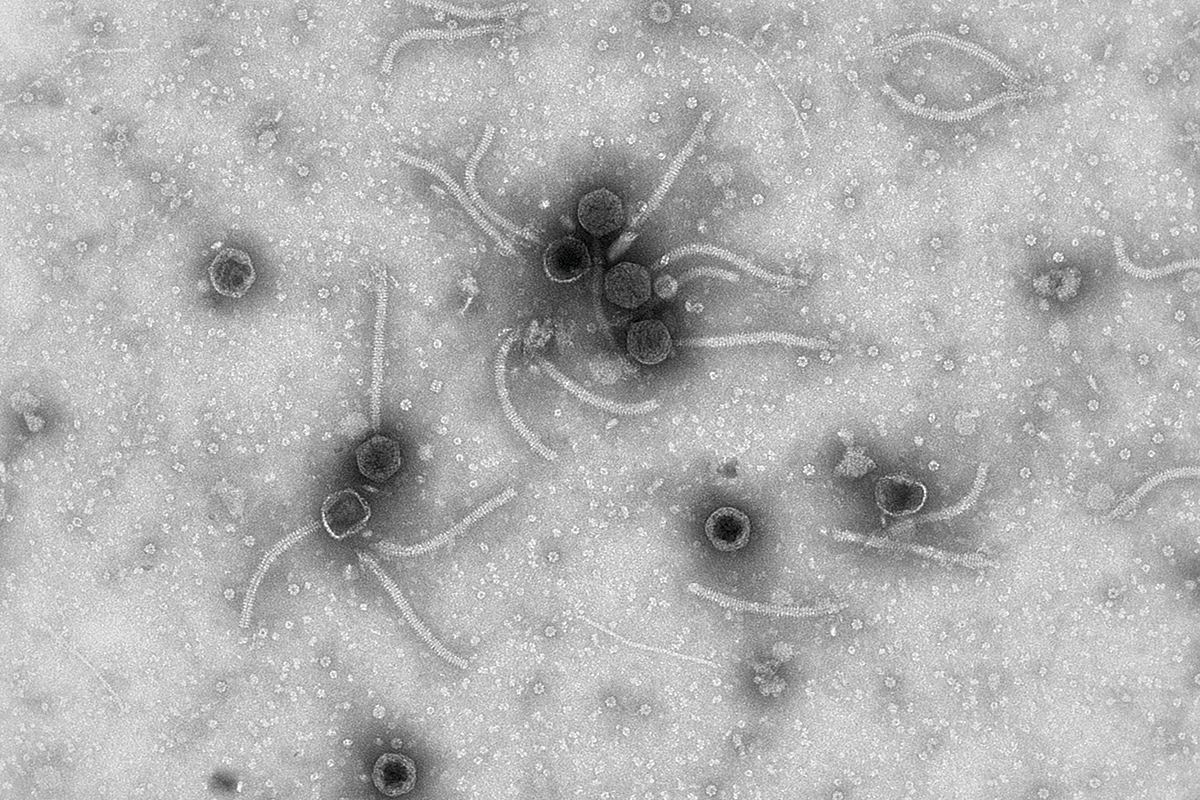
Figure 1. The images on this spread are examples of micrographs of phages, bacterial viruses, in which we can observe their morphological diversity. These micrographs were obtained at the Environmental and Biomedical Virology Laboratory of the Institute of Integrative Systems Biology (UV-CSIC). / Pilar Domingo-Calap
Bacteriophages, commonly known as phages, are the most abundant viruses in the biosphere. It is estimated that there are 1031 viral particles (that is, ten times more viruses than bacteria), and that for every bacterium, there is at least one phage capable of infecting it. These numbers reflect the importance of phages in ecosystems, as well as their enormous, as yet unknown, diversity. Phages’ genomes can be RNA or DNA, single- or double-stranded, circular or linear, segmented or unsegmented. In addition, a large number of morphologies have been observed, and the number keeps growing as new phages are discovered every year (Figure 1).
Depending on their life cycle, phages can be divided into lytic and temperate phages (although other alternative cycles have been described). The former destroy the host bacterium in a process known as lysis, which involves the rupture of the cell membrane and the release of the virus into the extracellular environment. Temperate phages incorporate their genome into the host genome and remain dormant in the form of prophages, and play a key role in horizontal gene transfer, i.e., the transmission of genetic information between organisms that does not occur through vertical transmission.
The discovery of phages and their first applications
Phages were independently discovered more than a century ago by the bacteriologist Frederick Twort (1915) and the microbiologist Félix d’Hérelle (1917). Both were looking for an explanation to the halo they sometimes observed when growing bacterial cultures on agar plates (Figure 2). However, D’Hérelle was the one to propose that the observed halo was the result of bacterial lysis caused by another micro-organism, which he called a bacteriophage. D’Hérelle also pioneered the use of phages as a treatment for children with dysentery in France and cholera in India, thus marking the beginning of phage therapy. Soon after, due to Alexander Fleming’s discovery of penicillin in 1928 and after World War II, the use of broad-spectrum antibiotics became widespread in Western countries and phage therapy was forgotten. Meanwhile, in Eastern countries, where antibiotics were largely unavailable, phages continued to be used as a treatment for bacterial infections, with worldwide reference centres that continue to use these therapies on a routine basis even today.
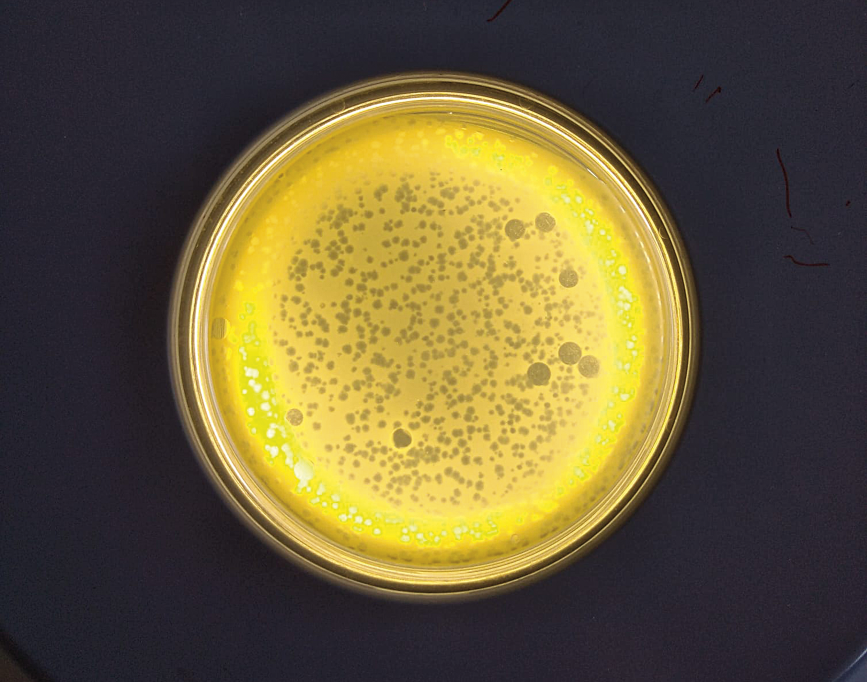
Figure 2. Examples of Petri dishes with bacterial cultures on semi-solid agar, showing the presence of lysis halos, which reflect the action of the phages lysing the bacteria growing on the plate. The one on the right contains two different phages, one that produces large halos and one that produces small ones. Lysis involves the rupture of the bacterial cell membrane and the release of the virus into the extracellular medium. Images taken at the Environmental and Biomedical Virology Laboratory of the Institute of Integrative Systems Biology (UV-CSIC). / Amanda Martínez
To date, Western countries lack specific legislation and clinical trials recognised and validated by regulatory agencies for the use of phages in patients. They are only allowed in exceptional cases and for compassionate use. However, scientists and physicians from all over the world are requesting regulatory agencies to include phage therapy as a recognised treatment. In our country, from the Spanish Network of Bacteriophages and Transducer Elements (FAGOMA), we are initiating the process through the Spanish Agency of Medicines and Medical Devices (AEMPS) to facilitate recognition of this therapy as an effective treatment against multidrug-resistant bacteria.
Phages in basic research
Phages have been widely used to understand fundamental biological processes such as the replication, transcription, and translation of nucleic acids. Notably, a large number of phage-derived products such as ligases and polymerases are commonly used as reagents in research. In Spain, the patent for the nucleic acid amplification method based on the DNA polymerase obtained from phage phi29 – discovered by Margarita Salas – continues to be the most profitable that the CSIC has ever presented. In fact, this technology is regarded as one of the great milestones in the history of science in our country. Recent Nobel Prizes in Chemistry have also been awarded for phage-based research. In 2018, Frances Arnold, George P. Smith, and Gregory Winter were awarded the prize for developing a method that used phages as vectors for the cloning or expression of genes, known as phage display, which facilitates the directed evolution of enzymes and antibodies. Using this technique, proteins of interest can be combined on the outside of a phage to create protein libraries that can be useful in various fields such as the development of antibodies, the study of protein-protein interactions, or the production of antivirals or antitumoural drugs, among many other applications. Recently, in 2020, Emmanuelle Charpentier and Jennifer Doudna were awarded the Nobel prize for their use of the bacterial immune system against phages as «genetic scissors», known as CRISPR-Cas9. This tool allows changes to be made to the DNA of any organism with high precision. These are just a few examples of the use of phages as experimental models in basic science and their potential applications today.
Phages in biomedicine
Phages are considered very promising tools in the field of biomedicine for their utility in the prevention, diagnosis, and treatment of bacterial diseases. Although their best known use is phage therapy, there are other applications of great interest such as biosensors, anti-tumour therapies, phage-based vaccines, or vehicles directed against specific targets in the field of nanomedicine. Thus, the potential of phages as biomedical tools is growing, thanks to their easy manipulation, high specificity, and low production cost. However, there are still weaknesses, such as the need to reassess the ethical aspects and regulatory protocols for their use in biomedicine. Here, we will detail some of the applications of phages in biomedicine, with the aim of raising awareness about their potential from different approaches.
Phages in prevention
The skin, mucous membranes, or the digestive tract are examples of compartments in which bacteria, viruses, and other micro-organisms that form our microbiome live. Phages and bacteria constantly co-evolve and keep the microbiome in balance. In this sense, phages can be used as «probiotics» to control bacterial communities and prevent diseases associated with inappropriate bacterial growth. Again, it should be noted that phages are highly specific to the bacteria they are able to recognise – thanks to specific receptors – and this is key to controlling bacterial strains of interest, as off-target bacteria remain unchanged and maintain homeostasis (Figure 3). Phages can therefore be considered microbiota modulators and tools, as they are part of the natural environment of the microbiome.
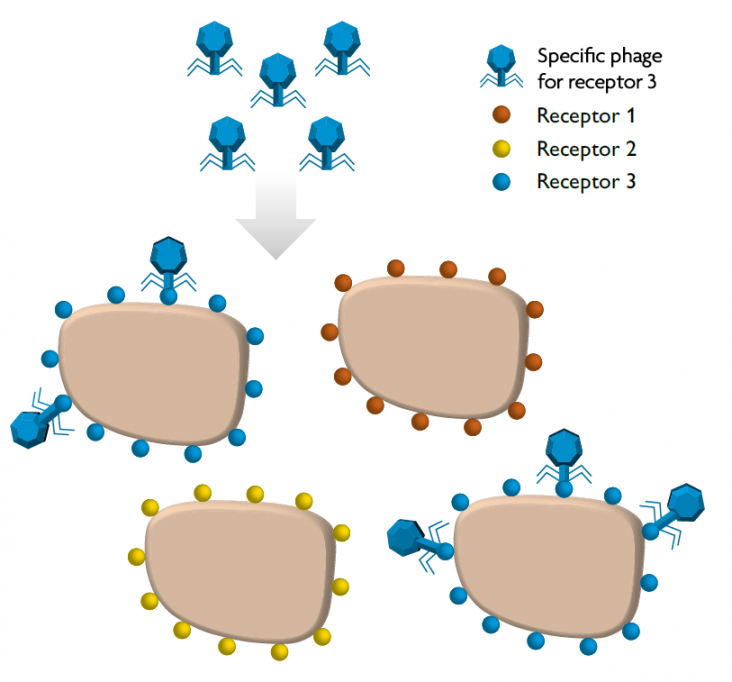
Figure 3. Due to their high specificity, phages can only recognise cells that express specific receptors. In the figure, we can see this process of recognition and binding of a given phage to a target bacterium. This functioning prevents the infection of other bacteria and potential side effects on the microbiota. / Adapted from Pilar Domingo-Calap
Another use of phages in disease prevention is the creation of phage-derived vaccines. Phage display-based engineering can produce specific vaccines against other pathogens such as fungi or even other viruses (Smith, 2019). Yet another application is to generate phage libraries for the production of recombinant antibodies. In this case, the antibodies obtained can be used as prophylactic agents, but also as diagnostic, bacterial typing, or therapeutic tools. Phages can also combat bacterial biofilms. In hospital environments, these biofilms form on medical devices, catheters, prostheses, or other surfaces, and are often composed of antibiotic-resistant nosocomial bacteria. Phages replicate in situ and, thanks to enzymes encoded in their genomes, can actively destroy these biofilms, penetrating and eliminating them very efficiently. Therefore, phages could be used to process medical devices or surfaces prior to use. It is worth noting that some phage-derived enzymes such as lysins or holins also have great potential to fight bacteria (Ferriol-González & Domingo-Calap, 2020).
Phages in diagnosis
As we have mentioned, phages can also be used for bacterial detection and typing, a quick and inexpensive solution to a sometimes laborious and difficult task. If we have a phage library capable of recognising a set of bacteria of interest, we can easily determine which bacteria are causing an infection. In this sense, phage-based bioimaging and biodetection is considered to be of particular relevance in biotechnology. In addition, it is possible to link specific markers in phage genomes to detect by imaging or sensors where these phages spread and to perform on-site tagging, which can help in the diagnosis of bacterial diseases and their specific localisation in the patient, because phages are very sensitive and can detect bacteria in very low quantities (Van der Merwe et al., 2014).
Phages as a treatment
The overuse of antibiotics has led to the emergence of multidrug-resistant bacteria, also known as superbugs. Unfortunately, today there are bacterial strains that cannot be treated with any of the available antibiotics. This problem has become a global threat and requires the development of alternative treatments. In fact, it is predicted that by 2050 there will be about ten million deaths per year related to resistant superbugs, more than cancer-related deaths (Kraker et al., 2016).
In recent years, the first clinical trials in phage therapy have started in Europe, and recently, because of the SARS-CoV-2 pandemic, the US has allowed the use of phages in COVID-19 patients in intensive care. The main reason is that half of the deaths in intensive care units were due to secondary infections caused by nosocomial superbugs causing pneumonia and sepsis. In particular, Acinetobacter baumannii, Staphylococcus aureus, and Pseudomonas aeruginosa have been successfully treated with phages in such patients. Thus, although the lack of legislation on phages hinders their use in clinical practice, the number of people treated with them is gradually increasing. Apart from COVID19 , patients themselves (and, in some cases, the medical staff) are currently contacting phage researchers directly to try to obtain specific viruses capable of controlling the target bacterial infections. In fact, patients with untreated superbug infections have made calls for help via social media to try phages as a therapeutic alternative. Fortunately, some of them have been successfully treated, opening the door to phage therapy as a treatment against pathogenic bacteria (Kortright et al., 2019). This process is still slow and complex, but more and more laboratories are getting involved and generating phage libraries to speed up the process.
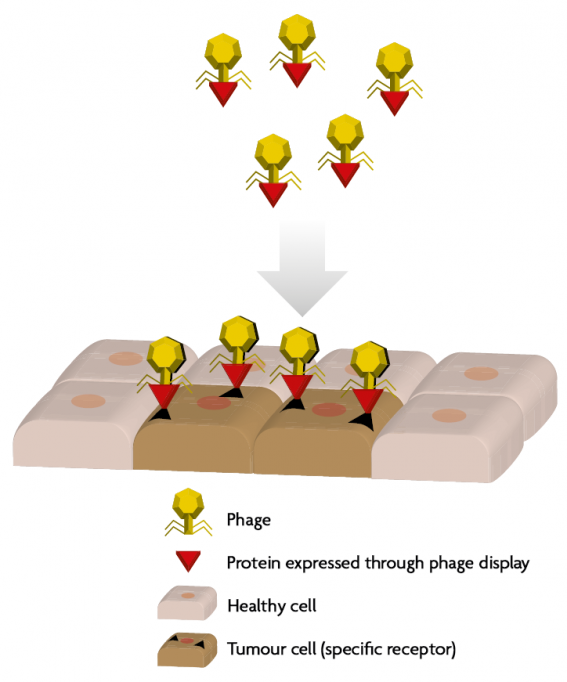
Figure 4. Genetically modified phages expressing exogenous proteins to recognise eukaryotic cells of interest, e.g., tumour cells. These cells express specific receptors that can be recognised thanks to the new proteins expressed in the phages (through genetic modification or phage display). Once the tumour cell is recognised, the phage can either act as a vehicle for anti-tumour molecules or induce cell death. / Adapted from Pilar Domingo-Calap
In contrast to broad-spectrum antibiotics, as discussed above, phages are highly specific, which is of particular interest to reduce side effects, since the microbiome will remain in balance and only the target bacteria will be destroyed. In the case of phage therapy, it is important to use lytic phages capable of rapidly and efficiently lysing bacteria. Notably, the use of therapies combining phages and antibiotics has also been proposed, as they have been found to have a synergistic effect.
Another therapeutic application of phages is, again, through phage display. It is possible to genetically modify phages to target cells of interest, including eukaryotic cells. Thus, phages can be used as vehicles to deliver genes or drugs to the desired cells, which is of particular interest in oncology, as phages can be engineered against tumour cells (Figure 4).
Conclusions
The use of phages in biomedicine is very diverse and allows the problem of resistant bacteria to be tackled from many perspectives (Domingo-Calap & Delgado-Martínez, 2018). These promising tools also have certain advantages over antibiotics, such as their low production cost, which would allow them to be used all over the world, even in low-income countries. On the other hand, just as antibiotic resistance has emerged, bacteria can also develop resistance to phages. However, unlike antibiotics, which are stable chemical molecules, phages evolve and adapt, and can counteract resistance. Moreover, it is possible to reduce the emergence of resistance through the directed evolution of phages in the laboratory. Another solution is the use of phage cocktails, in which the emergence of resistance can be significantly reduced.
However, despite their great versatility and very promising results, there are still weaknesses for their implementation in clinical practice. The main problems are the lack of legislation and their acceptance by the general public. Phages are viruses and their use may be hindered by ethical concerns. This is why regulatory protocols are necessary to reassure the public. Clinical trials and testing must follow (as yet non-existent) regulations to prove the safety of using phages in humans. In this regard, it is important to note that phages are incapable of infecting eukaryotic cells (unless genetically modified, as mentioned above), which makes them biologically and ecologically safe. It is also worth noting that, to date, research does not suggest that phages are recognised by the immune system, so there would be no specific response to them, which facilitates their administration, including intravenous delivery.
The enormous diversity of phages is also of particular interest at the ecological level, with new species being discovered and an unexplored biodiversity that may lead to new applications. In addition, new technologies linked to phages will help to create better tools in clinical practice. An interesting example is nanotechnology, which can improve the stability of phages, or help in their delivery or in the creation of biosensors. Multidisciplinary approaches in the field of biomedicine help us to explore new avenues and propose new personalised treatments with faster response and fewer side effects.
Finally, it should be noted that the use of phages is not limited to clinical use, but can have applications in many other fields such as agriculture and livestock farming. The same principles that we have been discussing can be applied to phytopathogenic bacteria or to bacteria that cause diseases in animals. This type of approach is also starting to be implemented in the agri-food industry, for example, in the United States and in some EU countries, where the use of phages is allowed for the control of pathogenic bacteria both in the food production chain and in the final product (Moye et al., 2018). More specifically, there are commercial phage preparations against bacteria such as Escherichia coli, Salmonella sp., and Listeria monocytogenes. Their commercialisation as biocontrol tools in food safety is proving very successful, both in the disinfection of industrial surfaces and in food decontamination.
Phages are very promising tools in the fight against resistant bacteria and, therefore, it is necessary to support this field of study in order to produce new data that favour their regulation and routine use. Public outreach and awareness-raising is also of great importance in order to reach the general public, to publicise new therapies for the control of pathogenic bacteria and to address the problem in an ecologically safe and sustainable way. Because not all viruses are bad. There are good viruses that can help us control the next potential pandemic: superbugs.
References
Domingo-Calap, P., & Delgado-Martínez, J. (2018). Bacteriophages: Protagonists of a post-antibiotic era. Antibiotics, 7(3), 66. https://doi.org/10.3390/antibiotics7030066
Ferriol-González, C., & Domingo-Calap, P. (2020). Phages for biofilm removal. Antibiotics, 9(5), 268. https://doi.org/10.3390/antibiotics9050268K
Kortright, K. E., Chan, B. K., Koff, J. L., & Turner, P. E. (2019). Phage therapy: A renewed approach to combat antibiotic-resistant bacteria. Cell Host & Microbe, 25(2), 219–232. https://doi.org/10.1016/j.chom.2019.01.014
Kraker, M. E. A. de, Stewardson, A. J., & Harbarth, S. (2016). Will 10 million people die a year due to antimicrobial resistance by 2050? PLOS Medicine, 13(11), e1002184. https://doi.org/10.1371/journal.pmed.1002184
Moye, Z. D., Woolston, J., & Sulakvelidze, A. (2018). Bacteriophage applications for food production and processing. Viruses, 10(4), 205. https://doi.org/10.3390/v10040205
Smith, G. P. (2019). Phage display: Simple evolution in a petri dish (Nobel lecture). Angewandte Chemie International Edition, 58(41), 14428–14437. https://doi.org/10.1002/anie.201908308
Van der Merwe, R. G., van Helden, P. D., Warren, R. M., Sampson, S. L., & van Pittius, N. C. G. (2014). Phage-based detection of bacterial pathogens. Analyst, 139(11), 2617–2626. https://doi.org/10.1039/C4AN00208C