A brief history of palaeogenomics
How a young discipline revolutionised the study of the past

In only a few short years, the ancient DNA field has transformed from an anecdotal and artisanal discipline into one of the most dynamic current scientific fields, generating massive genomic data from hundreds of past individuals. These include extinct hominins such as Neanderthals and Denisovans and prehistoric humans, and have provided information about the recent settlement of the continents. The field of palaeogenomics gives direct space and time information about the adaptive and demographic aspects of human populations and reveals complex patterns of past migrations that can help us to understand our current diversity. The development of this discipline is a unique opportunity to establish partnerships with archaeologists and anthropologists and to build up a multidisciplinary approach to the study of the past.
Keywords: human evolution, Neanderthals, genomics, prehistory.
The palaeogenomics (ancient DNA) field can be defined as the recovery and analysis of genetic material from the biological remains of the past and has become a powerful scientific field that provides direct information about the evolutionary process through space and time. Palaeogenomic findings, which include complete genomes of extinct hominids and modern humans from the past 50,000 years, have revolutionised our knowledge of human evolution, sometimes even ending archaeological and anthropological debates that had lasted for over a century. Moreover, this revolution has only just begun; soon there will be thousands of ancient genomes within the reach of all researchers.
«Over time, DNA sequences are fragmented in smaller and smaller pieces until they are impossible to attribute with certainty to any particular organism»
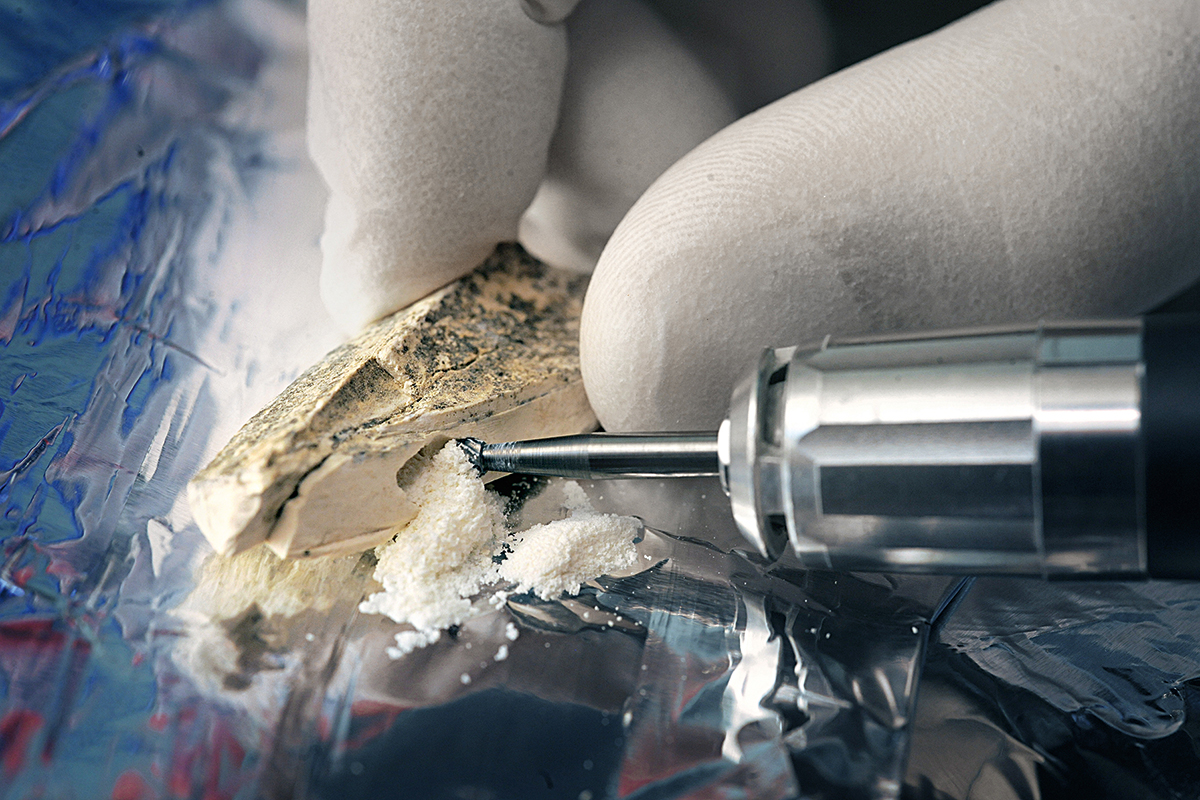
One of the specific features of the field is that this type of analysis requires the use of samples which potentially contain preserved DNA and that these are often unique; it is important to remember that, even though this technique only requires minimal quantities of skeletal material (either teeth or bone), it is still destructive. The state of conservation of these samples, whether they come from museums or directly from excavation sites, strongly depends on the climate conditions (essentially, the thermal conditions) in which it was preserved. The cooler the environment, the higher our chance of being able to use its DNA to go back in time. Under ideal conditions, like those of Siberian frozen soil, we can expect to be able to trace a history of up to a million years. For example, the genome of a Pleistocene equid was found to date back to between 560,000 and 780,000 years). In temperate conditions like those in most of Europe, the current antiquity record is about 430,000 years, but it usually goes back only a few tens of thousands of years. Finally, in very hot climates (where, unfortunately, many key evolutionary processes of the human lineage took place), we would be lucky to be able to rescue even a few thousand years. It is worth noting that these time limits will not improve with any technological advances because, over time, DNA sequences are fragmented into smaller and smaller pieces until they become impossible to attribute with certainty to any particular organism.
Another of the differential characteristics of this field is the fact that it developed together with technical breakthroughs, and especially with new massive sequencing platforms created in the last ten years, which have turned this anecdotal and artisanal scientific field into an almost conventional discipline with some features of mass production. Very few laboratories have managed to survive the transformation from one methodology to the other. That is, to move from experimental to computational techniques, from analysing small DNA fragments to using entire genomes, and from working with one or a few samples to working with hundreds; as a result, the field is currently dominated by perhaps a dozen laboratories, most of which, interestingly, are based in Europe. Although some assumed that new technologies would democratise the discipline and make it more affordable, in reality it has become more technical and expensive, and the funding resources – which are more scarce given the current financial crisis – have increasingly concentrated in a few reference centres.
«The field of ancient DNA is considered to have started officially in 1984, with the recovery of DNA sequences from a quagga»
But the collaboration between geneticists, archaeologists, and anthropologists did improve, to finally reach a truly multidisciplinary view of the study of the past. This occurred mainly because new genomic data allowed us to explore more specific questions of interest than previously, like how to determine genetic sex or the kinship relationships between different individuals in the same site. Additionally, we can now analyse individuals closer to the present, in periods about which we have abundant historical and archaeological information.
The heroic era (1984-1997)
The field of ancient DNA is considered to have started officially in 1984, with the recovery of DNA sequences from a quagga, a South African equid extinct since the late nineteenth century, thanks to a conserved naturalised specimen (Higuchi, Bowman, Freiberger, Ryder, & Wilson, 1984). The following year, Svante Pääbo, the historical leader of the discipline, published data from the first DNA recovery from human remains, specifically from an Egyptian mummy (Pääbo, 1985). Both studies used the bacteria cloning technique to recover small fragments of the material; however, the procedure is quite inefficient because it is unspecific. We must also consider the fact that the polymerase chain reaction technique (or PCR), which dominated molecular genetics in the following twenty years, had not yet been invented. While the validity of the first study was later confirmed, the second is now commonly believed to be the result of modern DNA contamination.
«Palaeogenomics developed together with technical breakthroughs, and especially with new massive sequencing platforms»
The development of PCR, which allows experts to recover specific DNA fragments for sequencing, and the discovery that genetic material survived also in skeletal material and not only in mummies, led to the constant diversification and increase in ancient DNA studies. Very soon, however, it was noted that the technique would favour the recovery of external or contaminant DNA from people who had handled the studied remains, including archaeologists or museum curators. This led to some erroneous results and to the development of more and more sophisticated procedures to maintain laboratories as clean and isolated as possible. Some groups were not careful enough and published implausible reports of DNA recovery from remains that were tens of millions of years old, including DNA from Miocene tree leaves, from insects conserved in amber, or from Cretaceous dinosaur bones. It is worth noting that competition between the two most important scientific journals, Science and Nature, made it easier for these studies, which lacked proper verification, to be published in one or the other.
This situation caused the field to lose some prestige, but at the same time it triggered the adoption of several control measures that any group intending to publish their work had to follow, the key one being independent replication of the results by a different laboratory (Cooper & Poinar, 2000). This procedure helped establish a series of collaborative links between laboratories, but also limited the number of laboratories with the necessary technical level.
All these efforts crystallised in the first recovery of Neanderthal mitochondrial DNA (specifically from the original skeleton at the Feldhofer cave, in the German valley of Neander, dating back around 40,000 years). The work was led by Svante Pääbo and illustrated the cover of the journal Cell in 1997 (Krings et al., 1997). The problem of contamination was dismissed because the mitochondrial DNA sequence of the Neanderthal was very different from that of modern humans. At the same time, this meant that modern humans and Neanderthals belonged to different evolutionary lineages, at least with respect to mitochondrial DNA.
In the following years, several mitochondrial DNA sequences from other Neanderthals slowly accumulated in the literature, but was still limited by the technology available at the time. From among these, the first sequence of an Iberian Neanderthal, from the cave of El Sidrón (Asturias), published in 2005 was especially noteworthy. This work also proved the low genetic diversity of Neanderthals, which indicated that its population size was very small.
Consolidation work (1997-2010)
In the following ten years, the field started to consolidate itself and proved its status, beyond the anecdotal, as a useful scientific tool. It also attempted to understand methodological mechanisms such as post mortem chemical damage patterns and the fragmentation pattern of DNA strands over time (Hofreiter, Serre, Poinar, Kuch, & Pääbo, 2001). Since contamination problems were less important when working with extinct animals (the probability of contaminating a mammoth bone with elephant DNA is nowhere near the probability of contaminating a human bone with modern human DNA), palaeogeneticists focused mainly on creating phylogenies that allowed us to understand the evolutionary affinities of extinct species over the last thousand years. These species included paradigmatic animals like cave bears, woolly rhinoceros, ground sloths, thylacines, mammoths, New Zealand moas, and Myotragus, the latter being a caprine endemism of the Balearic islands, which became extinct about 4,500 years ago because of the arrival of the first humans to the islands.
All this work recovered mitochondrial DNA, a small genome located within cell mitochondria. It has the advantage of being represented in a very high number of copies compared to nuclear DNA. However, at the same time, technical limitations implied that the discipline would never be able to turn towards more extensive and informative nuclear genomics. The most interesting breakthrough came in 2001, when the entire mitochondrial genome of two moa species was recovered. It was an enormous task, consisting of overlapping dozens of small fragments recovered by PCR, which no one has dared repeat since (Cooper et al., 2001).
«The realisation that human remains were already contaminated by the time they reached the ancient DNA laboratories led to the development of anticontamination protocols in the excavation sites»
The realisation that human remains had already been contaminated by the time they reached the palaeogenomics laboratories led to the search for fresher samples and to the development of anticontamination protocols at excavation sites. In this sense, the recovery, in controlled conditions and using sterile suits, of Neanderthal remains at the site of El Sidrón, became a scientific benchmark for many human evolution and archaeology books.
All the aforementioned allowed experts to start exploring fragments of nuclear genes, which in the case of the Neanderthals, proved the existence of two mutations which are shared with modern humans: one in a key language-related gene and the other, a specific mutation that caused reddish hair like that of modern red-haired people. In this second study, the researchers used functional genomics techniques to obtain pigment cells in vitro that expressed the Neanderthal protein in their membranes (Lalueza-Fox et al., 2007). This signified the first time that ancient DNA studies abandoned its purely phylogenetic approach to explore the phenotypic and adaptive aspects of extinct humans.

The genomic revolution (since 2010)
At the end of the first decade of this century, no one could have imagined that ancient DNA would become the most revolutionary scientific field in the study of past human beings, or that it would be applied almost to an industrial scale. The change came thanks to new massive sequencing technologies, known as second generation sequencing technologies, which emerged for the first time in 2005 and became popular a few years later. Complete ancient genomes have been obtained thanks to these technological platforms, sometimes with a higher sequence quality than the ones published using current samples. In addition, they have allowed us to understand the basic aspects of DNA fragmentation, which occurs because of a chemical process which leaves a distinctive signal at the ends of ancient sequences, a signal that cannot be found in modern contaminants. This way, the bane of contamination was left behind: using this chemical pattern, scientists could even work with contaminated samples.
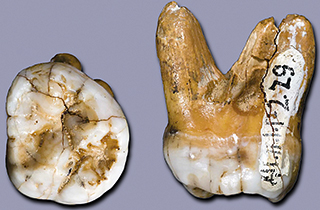
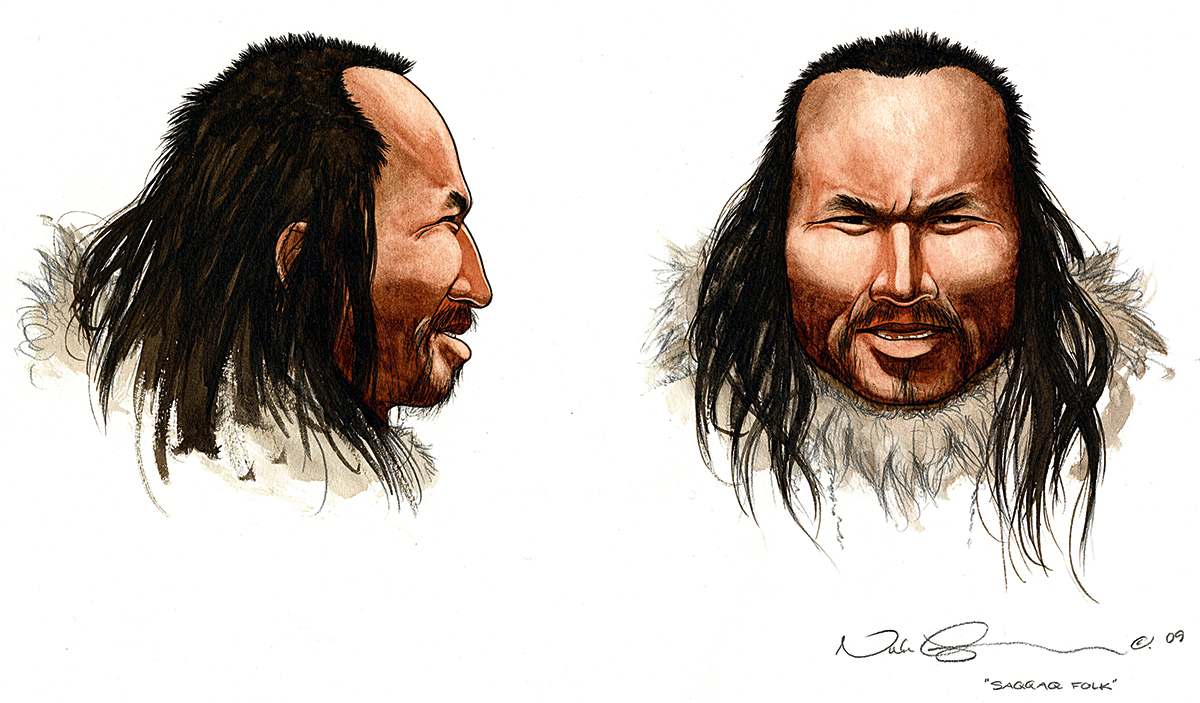
In 2010, thanks to all these advances, several publications arrived: the first ancient genome of a modern human, the Saqqaq man, a 5,500-year-old Palaeo-Eskimo found in Greenland (Rasmussen et al., 2010), the first Neanderthal genome, using three different individuals at the Croatian site of Vindija (Green et al., 2010), and an Asian hominin from the Denisova cave in Russia, which has not yet been taxonomically defined (Reich et al., 2010). The two latter genomes represented a paradigm change in the understanding of the human evolutionary process, because they delimited a list of genes that presented amino acid differences between us and other extinct human lineages – and, therefore, of genes that are likely the basis of what makes us different as a species – and because they provided direct evidence of several hybridisation processes between modern humans, Neanderthals, and Denisovans. Such crossbreeding is not reflected in mitochondrial DNA because of its transmission exclusively through maternal lines; therefore, the mitochondrial genomes of Neanderthals and modern humans remain different. However, we now know that Neanderthals contributed around 2% to modern non-African human DNA, and the Denisovans provided around 4% of the Australasian aborigine genome. Some of the ancient genes acquired by modern humans were later selected because they represented an adaptive advantage in the newly colonised environments, in traits such as protection from certain diseases or height adaptations. At the same time, some of them have led to negative effects in current populations, such as genetic variants involved in cardiovascular disorders or in diabetes. This information represents a paradigm change in our conceptualisation of human evolution, which should be now be understood as a recent exodus from Africa but with many hybridisation episodes with other hominins. These processes also explained the difficulties of defining species using only the fossil record, because it seems clear that this mixing might have happened much earlier, millions of years ago.
«Second generation sequencing technologies allowed us to know about basic aspects of DNA fragmentation. The bane of contamination was left behind»
In 2014, new work explored the limits of the new techniques and shook the hominin tree, recovering a complete mitochondrial genome using remains from Sima de los Huesos at the Atapuerca site, dating back about 430,000 years. The quality of the record was phenomenal, which is partly explained by the unique conservation conditions of the interior of the cave (Meyer et al., 2014). The mitochondrial DNA phylogeny indicated that the hominin was related to Denisovans, even though the physical features of the Sima de los Huesos skulls had been interpreted as ancestors of the Neanderthals. Even so, the recovery of small nuclear genome fragments from other individuals over the next couple years (Meyer et al., 2016), confirmed the affinity with Neanderthals. Once again, the discrepancy between mitochondrial and nuclear data provided evidence that a complex migration and hybridisation pattern had occurred during the Pleistocene within the evolutionary history of these hominins.
«Palaeogenomics has also transformed European prehistory. A series of studies allowed us to explore the evolutionary changes in key characteristics»
The next field that was profoundly transformed by advances in palaeogenomics was European prehistory. Starting in 2012, a series of studies allowed us to explore the evolutionary changes in key characteristics such as pigmentation, diet, metabolism, or susceptibility to several infectious diseases, associated with the arrival of agriculture and to subsequent migrations from central Asia. The first Mesolithic genome (from La Braña, in León) was published in Nature in 2014 (Olalde et al., 2014), and only three years later we already had data from hundreds of specimens from the Mesolithic, Neolithic, Copper Age, and Bronze Age, and even later eras (Haak et al., 2015; Lazaridis et al., 2014). These studies determined that current European populations are the result of three genetic components which overlap in different proportions depending on the population: Mesolithic hunter-gatherers, Neolithic farmers from the Middle East, and nomads, the so-called Yamnaya, from the steppes who came from the East in the late Neolithic (Haak et al., 2015). This new transversal vision of European ancestry explains the peculiarities and similarities between populations within the same continent and in relation to neighbouring territories. Southernmost populations have fewer components from the steppes and westernmost ones have greater Mesolithic and lower Neolithic components. The expansion of the steppes component was also connected to the spread of Indo-European languages first and Celtic languages later, and is correlated with an exceptional increase in one lineage on the Y chromosome (R1), which is predominant in modern Europe. Thus, this new information provides a view that interrelates genetics, demography, social structure, and culture.
Genomic data have also been recovered from dozens of European Palaeolithic remains. It has revealed new hybridisations with the last Neanderthals (like in the case of Oase, in Romania) and how a complex series of migrations and population replacements took place over the last 45,000 years. It is worth noting that Palaeolithic settlers such as Oase have not left any genetic traces in current-day Europeans because the Mesolithic component was derived from contributions from migrants from outside of Europe about 14,000 years ago, after the last glacial peak. Therefore, we do not descend from Upper Palaeolithic Europeans (Fu et al., 2016).
«In a few years we will have hundreds, maybe thousands, of ancient genomes, especially from temperate areas like Europe»
The palaeogenomic revolution did not only impact the study of past humans. It also revolutionised our understanding of their diseases. Pathogen evolution models the adaptation of human populations, given that we are the descendants of ancestors that survived past epidemics. The bacterium that caused the plague was recovered from the remains of individuals who died during several historical outbreaks, such as the Black Death in the Middle Ages (which reached Europe in 1348) or the Plague of Justinian, with the first declared outbreak, between 541 and 543. But a recent analysis also detected the pathogen in the migrants who came to Europe from central Asia’s steppes, and who radically transformed the genetic makeup of the continent during the Bronze Age. Local Neolithic populations had never been in contact with this disease before, therefore, the mortality caused by this recently discovered prehistoric epidemic could also explain the great demographic change related to the arrival of nomads from the steppes.

The future of palaeogenomics
In a few years we will have hundreds, maybe thousands, of ancient genomes, especially from temperate areas like Europe. But data from individuals from other continents will also be collected, including those from Africa, a continent whose only available information is the Mota genome, from Ethiopia, dating back about 4,000 years. The automated processing of these analyses will allow the quasi-industrial production of information, and palaeogenomics will be considered almost a service, just as radiocarbon dating is today. Surely, new data from very ancient hominins will appear (albeit limited to within the last hundreds of thousands of years), especially in little-explored areas with a favourable climate, such as Asia. Genome statistics interpretations will be used to reconstruct past migratory movements, but also to provide live knowledge about the evolutionary process, with complementary data like the temporal description of adaptive or demographic phenomena. Integrating all this wealth of information will lead to a new vision of the study of the past, which will become more global and interdisciplinary, and will close many scientific debates that seemed unresolvable only a few years ago.
REFERENCES
Cooper, A., Lalueza-Fox, C., Anderson, S., Rambaut, A., Austin, J., & Ward, R. (2001). Complete mitochondrial genome sequences of two extinct moas clarify ratite evolution. Nature, 409, 704–707. doi: 10.1038/35055536
Cooper, A., & Poinar, H. N. (2000). Ancient DNA: Do it right or not at all. Science, 289, 1139. doi: 10.1126/science.289.5482.1139b
Fu, Q., Posth, C., Hajdinjak, M., Petr, M., Mallick, S., Fernandes, D., ... Reich, D. (2016). The genetic history of Ice Age Europe. Nature, 534, 200–205. doi: 10.1038/nature17993
Green, R. E., Krause, J., Briggs, A. W., Maricic, T., Stenzel, U., Kircher, M., ... Pääbo, S. (2010). A draft sequence of the Neandertal genome. Science, 328, 710–722. doi: 10.1126/science.1188021
Haak, W., Lazaridis, I., Patterson, N., Rohland, N., Mallick, S., Llamas, B., ... Reich, D. (2015). Massive migration from the steppe was a source for Indo-European languages in Europe. Nature, 522, 207–211. doi: 10.1038/nature14317
Higuchi, R., Bowman, B., Freiberger, M., Ryder, O. A., & Wilson, A. C. (1984). DNA sequences from the quagga, an extinct member of the horse family. Nature, 312, 282–284. doi: 10.1038/312282a0
Hofreiter, M., Serre, D., Poinar, H. N., Kuch, M., & Pääbo, S. (2001). Ancient DNA. Nature Reviews Genetics, 2, 353–359. doi: 10.1038/35072071
Krings, M., Stone, A., Schmitz, R. W., Krainitzki, H., Stoneking, M., & Pääbo, S. (1997). Neandertal DNA sequences and the origin of modern humans. Cell, 90, 19–30. doi: 10.1016/S0092-8674(00)80310-4
Lalueza-Fox, C., Römpler, H., Caramelli, D., Stäubert, C., Catalano, G., Hughes, D., ... Hofreiter, M. (2007). A melanocortin 1 receptor allele suggests varying pigmentation among Neanderthals. Science, 318, 1453–1455. doi: 10.1126/science.1147417
Lazaridis, I., Patterson, N., Mittnik, A., Renaud, G., Mallick, S., Kirsanow, K., ... Krause, J. (2014). Ancient human genomes suggest three ancestral populations for present-day Europeans. Nature, 513, 409–413. doi: 10.1038/nature13673
Meyer, M., Arsuaga, J. L., De Filippo, C., Nagel, S., Aximu-Petri, A., Nickel, B., ... Pääbo, S. (2016). Nuclear DNA sequences from the Middle Pleistocene Sima de los Huesos hominins. Nature, 531, 504–507. doi: 10.1038/nature17405
Meyer, M., Fu, Q., Aximu-Petri, A., Glocke, I., Nickel, B., Arsuaga, J. L., ... Pääbo, S. (2014). A mitochondrial genome sequence of a hominin from Sima de los Huesos. Nature, 505, 403–406. doi: 10.1038/nature12788
Olalde, I., Allentoft, M. E., Sánchez-Quinto, F., Santpere, G., Chiang, C. W. K., DeGiorgio, M., ... Lalueza-Fox, C. (2014). Derived immune and ancestral pigmentation alleles in a 7,000-year-old Mesolithic European. Nature, 507, 225–228. doi: 10.1038/nature12960
Pääbo, S. (1985). Molecular cloning of Ancient Egyptian mummy DNA. Nature, 314, 644–645. doi: 10.1038/314644a0
Rasmussen, M., Li, Y., Lindgreen, S., Pedersen, J. S., Albrechtsen, A., Moltke, I., ... Willerslev, E. (2010). Ancient human genome sequence of an extinct Palaeo-Eskimo. Nature, 463, 757–762. doi: 10.1038/nature08835
Reich, D., Green, R. E., Kircher, M., Krause, J., Patterson, N., Durand, E. Y., ... Pääbo, S. (2010). Genetic history of an archaic hominin group from Denisova Cave in Siberia. Nature, 468, 1053–1060. doi: 10.1038/nature09710