The cenancestor is defined as the last common ancestor of every currently living being. Its nature has been inferred from the identification of homologous genes between archaea, bacteria, and eukaryotic lineages. These inferences indicate that the cenancestor had a relatively modern protein translation system, similar in complexity to that of a current cell. However, the key enzymes for the replication of genetic material and for cell membrane biosynthesis are not homologous in bacteria, archaea, and eukaryotes. Here, we briefly review the history of the concept of the last universal common ancestor and the different hypotheses proposed for its biology.
Keywords: universal phylogeny, LUCA, horizontal transference, early evolution, common ancestor.
Evolutionary biologists are in the same logical predicament as historians and can only present arguments based on the assumption that, of all the plausible historical sequences, one is more likely to be a correct description of the past events than another.
Lynn Margulis, 1975, 29:21-38
The concept of homology and the universal tree of life
They say that a picture is worth a thousand words. In the theory of evolution, one of the most suggestive images can be found in a notebook written by Darwin before On the origin of species was published (Figure 1). In the book, we can appreciate how Darwin represents the origin of different species from a common ancestor in his drawing.
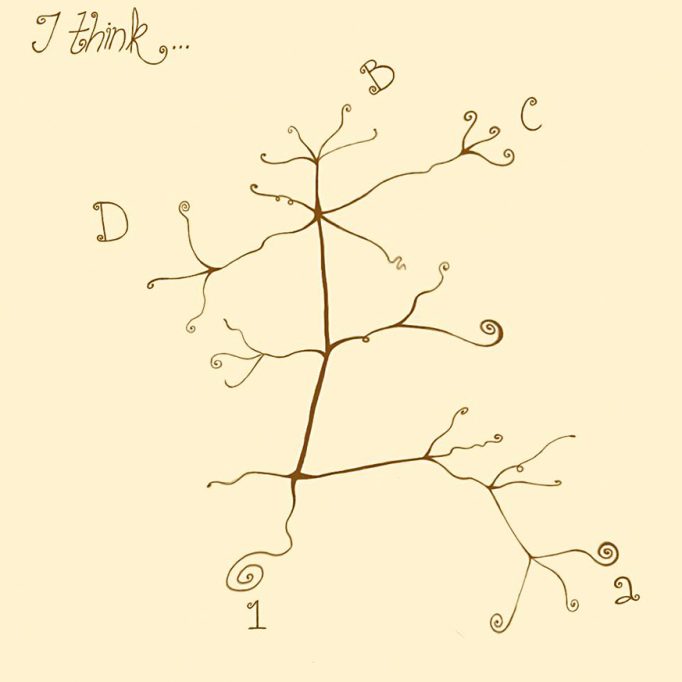
Figure 1. Darwin outlined the evolution of a group of species from an ancestral species in a notebook known as«Notebook B». / Drawn by Sofía Delaye Pascual
As we can see, Darwin’s theory of evolution suggests that the different species living today have evolved away (diversified) from each other from common ancestors. This apparently simple idea profoundly changed the way we understand living beings. For instance, when we compare the limbs of different mammals, we find that their bones are very similar to each other (Figure 2). According to Darwin, this similarity exists because those limbs evolved from a common ancestor; that is, they are homologous. In fact, Darwin wrote in On the origin (1859, p. 415):
What can be more curious, than that the hand of a man, formed for grasping, that of a mole for digging, the leg of the horse, the paddle of the porpoise, and the wing of the bat, should all be constructed on the same pattern […]?
The concept of homology allows us to deduce some of the features of the common ancestor for a certain group of organisms. For instance, in the case of the mammals in Figure 2, we can deduce that the common ancestor for all of them had these bones in the same relative positions. Darwin’s theory opens the possibility that all living beings are related through a great universal tree of life and evolved from a single common ancestor. Indeed, Darwin suggested in On the origin (1959, p. 455): «[…that] all the organic beings which have ever lived on this earth have descended from some one primordial form.»
However, Darwin, as cautious as ever in his scientific statements, never dared to represent the evolution of every living being in a single tree. Ernst Haeckel was the one of the first to draw a universal tree. As we can appreciate from Figure 3, Haeckel suggests that living beings diverge from a common trunk into plants, animals, and protists.
«Darwin’s theory opens the possibility that all living beings are related through a great universal tree of life and evolved from a single common ancestor»
Subsequently, during the first half of the twentieth century, Chatton (1938) and, even more clearly, Stanier and van Niel (1941) suggested that living beings should be classified in two main groups, i.e., prokaryotes and eukaryotes (Figure 4). The main features of eukaryotes are that they possess internal membranes, like the nuclear membrane and the endoplasmic reticulum, and that they divide by mitosis, while prokaryotes do not have internal membranes and divide by fission. In the words of Stanier and van Niel (1941, p. 464): «…the Monera kingdom, composed of microorganisms without true nuclei, plastids, and sexual reproduction.»
In 1969, the ecologist Robert Whittaker suggested that living beings should be classified into four great eukaryotic kingdoms (Plantae, Animalia, Protists, and Fungi) and one prokaryote kingdom (Monera). This classification scheme was adapted by Lynn Margulis and combined with the prokaryote/eukaryote division to accommodate the origin of mitochondria and chloroplasts, respectively, from alphaproteobacteria and cyanobacteria (Figure 5). However, the advent of molecular biology brought the issue of the nature of the last common ancestor to the foreground of research.
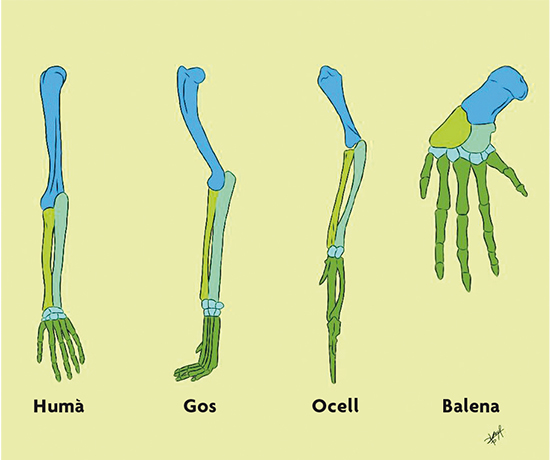
Figure 2. Limb homologues from different mammals. From left to right, human, dog, bird and whale. / Drawn by Sofía Delaye Pascual
The universal ancestor, history of the concept
In the middle of the twentieth century, Frederick Sanger developed the necessary techniques to understand the amino acid and nucleotide sequences that make up proteins and DNA, respectively. The information generated with these techniques allowed Emile Zuckerkandl and Linus Pauling to suggest that it was possible to reconstruct the history of living beings by comparing these biomolecules.
In 1977, using molecular techniques, one of the most surprising discoveries about the diversity of life on Earth was published. Carl Woese and George Fox discovered that, regarding the similarities and differences in the small subunit of the ribosomal RNA molecule (SSU rRNA), living beings are divided into three groups (Woese & Fox, 1977a). These three groups (or lineages) are: (a) eubacteria; (b) the eukaryotic nucleus-cytoplasm; and (c) a group of prokaryotes named archaebacteria. These three groups are currently known as Bacteria, Eukaryotes, and Archaea.
Based on this universal division, Woese and Fox suggested that there was a primitive entity in the divergence of these three cell lineages, when the relationship between genotype and phenotype had not yet evolved to its current form (Woese & Fox, 1977b, p. 1). They named this ancestral biological entity a progenote: «This primitive entity is called a progenote, to recognize the possibility that it had not yet completed evolving the link between genotype and phenotype». Some years later, Fitch and Upper (1987, p. 761), when studying the evolution of the genetic code, coined the term cenancestor, defined as: «the most recent common ancestor to all the organisms that are alive today».
«The advent of molecular biology brought the issue of the nature of the last common ancestor to the foreground of research»
It is important to note the differences between both concepts. The progenote implies a primitive state, while the concept of the cenancestor does not, at least not necessarily. The cenancestor, also known as the Last Universal Common Ancestor or LUCA, is the last common ancestor only of currently living beings, and it could be as simple as a progenote or as complex as a current cell.
Thanks to the development of DNA sequencing technology and to the accumulation of a vast quantity and diversity of sequences in databases, it was possible to start identifying the genes conserved in Bacteria, Archaea, and Eukaryotes (Figure 6). This enabled a simple methodology to infer the genes that were already present in the cenancestor genome. Based on this methodology, Lazcano, Fox and Oró (1992) suggested that the last common ancestor possessed genetic machinery similar to that of a current prokaryote cell.
However, the phylogeny proposed by Woese and Fox lacks a root, because it does not show whether any of the lineages is older than the other two. The first approach to identifying the root of the universal tree was independently made by two research groups using universally-conserved duplicate genes. Both analyses suggested that the root of the universal tree would be the bacteria branch (Figure 7). This reinforced the idea that the cenancestor was prokaryotic in nature.
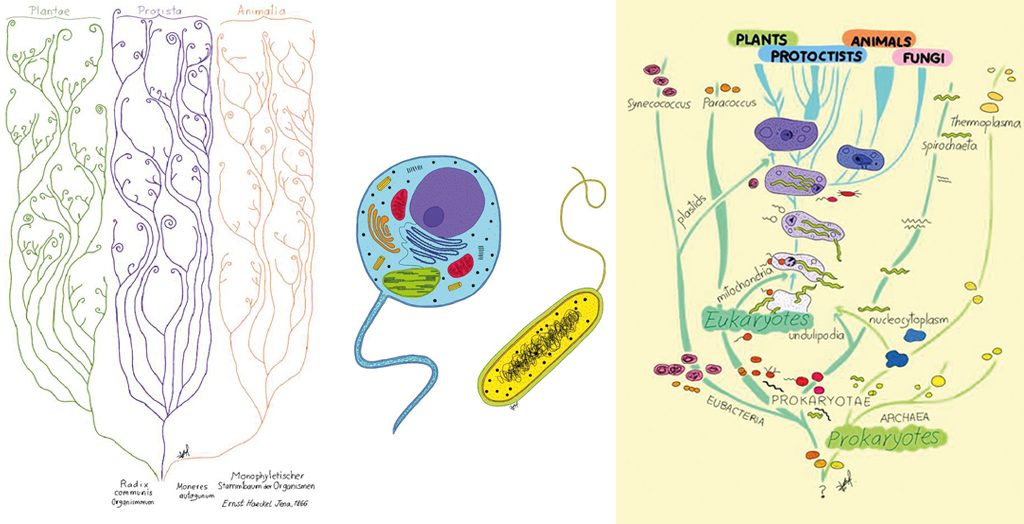
On the left, Figure 3, tree of life inspired by Generelle Morphologie der Organismen (1866), by Ernst Haeckel. In the middle, Figure 4, on a cellular level, we can classify all the living beings as prokaryotes and eukaryotes. On the right, Figure 5, the five kingdoms classification-scheme amended by Lynn Margulis (1996). In the picture, we can see the cytological division of all living beings into prokaryotes and eukaryotes. We can also observe how prokaryotes (Archaea and Bacteria) originate eukaryotes (Animals, Plants, Fungi, and Protists) through symbiogenesis. / Drawn by Sofía Delaye Pascual
RNA or DNA?
In 1996, Mushegian and Koonin compared the genomes of two bacteria, Haemophilus influenzae and Mycoplasma genitalium, in order to propose the minimal and sufficient set of genes necessary to support cellular life. They also looked for genes that were homologues to H. influenzae and M. genitalium in Archaea and Eukaryotes, in order to identify universally conserved genes and to deduce the nature of the last common ancestor. As they did not find homologous genes for the enzyme in charge of DNA replication in all three cell lineages, Mushegian and Koonin proposed that the last common ancestor for all living beings had an RNA genome. This would mean that the last common ancestor would be simpler than any currently known cell. Nonetheless, further inferences have proven that the set of conserved genes is not consistent with the complexity level of a progenote.
Horizontal gene transfer and the tree of life
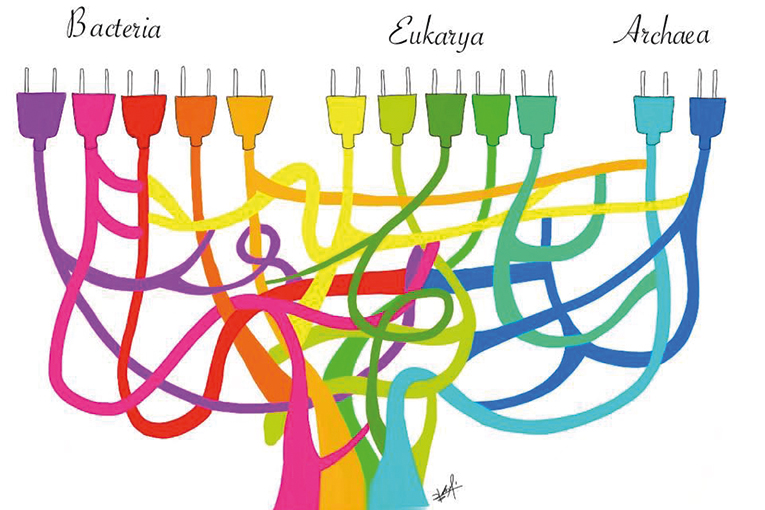
One of the main surprises coming from complete genome sequencing at the end of the nineties was the frequent lack of concordance in the phylogeny suggested for rRNA, where living beings group into Archaea, Bacteria, and Eukaryotes, with phylogenies derived from other genes preserved in these three cell lineages. While experts thought at the beginning that many of the discrepancies stemmed from methodological problems, subsequent studies proved that, at least among prokaryotes, an important proportion of phylogenetic incongruities were due to horizontal gene transfer (Figure 8).
«The cenancestor, also known as the Last Universal Common Ancestor or LUCA, is the last common ancestor only of currently living beings»
The discovery of an abundance of horizontal transfer events throughout the history of life on Earth posed a major problem for the reconstruction of the set of genes present in the last common ancestor. In fact, it was even suggested that during the early stages of the evolution of life on Earth, horizontal inheritance was the most common form (Woese, 1998).
New genes, lost genes, and the cenancestor
On the one hand, a genome can gain genes through horizontal transfer or through de novo creation. On the other hand, a gene that does not contribute to the survival of an organism anymore can get lost throughout evolution. As we have seen, the frequency of these events affects the reconstruction of gene content from the last common ancestor. In an attempt to infer the content of the cenancestor genes, taking into account the events of gene gain or loss throughout the history of life on Earth, Mirkin and colleagues (2003) developed an algorithm based on the argument of parsimony. According to their algorithm, the last common ancestor had around 572 genes. This set of genes was almost enough to code a coherent metabolism. However, there are two important absences: The first, as we said before, was the lack of DNA polymerase; the second great absence are two key enzymes in charge of synthesising cell membrane lipids.
The lack of these components is surprising if we take into account that they represent two of the three most important properties of living beings, that is: the separation of a living being from its environment by a plasma membrane and the legacy of genetic material based on the replication of a nucleic acid (Szathmáry, 2005). The other important feature of living beings, explored by Eörs Szathmáry and present in Mirkin’s reconstruction, was metabolism. The dispute over the nature of the genome of the last common ancestor and the nature of its membranes is still open in the scientific community (Peretó, López García, & Moreira, 2004; Poole et al., 2014).
The last common ancestor and viruses
Another unclear aspect about the nature of the cenancestor is whether or not viruses existed at the time and whether or not they could «infect» it. We still do not have conclusive evidence today of the co-existence of viruses and cells as old as the common ancestor. However, different hypotheses suggest not only that these entities coincided, but that viruses predate cells, or even that they played a central role in the very early stages of life, close to its origin (Agol, 2010).
«A gene that does not contribute to the survival of an organism anymore can get lost throughout evolution»
There are many unresolved questions concerning the origin and evolution of viruses. Nonetheless, many researchers consider, as do we, that the origin of these biological entities is polyphyletic. Therefore, it is very difficult, or even impossible, to prove their presence in the early stages of life due to their nature as «escaped» material from cellular organisms. However, this stance is far from having a wide consensus and faces a number of objections. For example, the fact that a few (important) protein structures, such as the jelly-roll of the capsid, are present in different virus types suggests a common origin. In addition, a small list of possibly homologous genes was proposed for several virus groups, those called hallmark genes (Koonin & Dolja, 2013), which added weight to the monophyletic proposal. If true, this information would be evidence of a common origin for all viruses, and suggests that such a viral ancestor (or ancestors) was contemporary, or at least, present at a similar time to the cellular ancestor.
It is important to note that the information we have now does not allow for discussion on the presence of viruses at times close to the cenancestor, and data cannot be extrapolated as evidence for earlier stages of life. Neither can we suggest that viruses played a central role in the transition from RNA-genome organisms to DNA-genome organisms, or even in the origin of life. Molecular methods and comparing sequences cannot provide information from pre-cellular stages.
Despite this fact, if the last common ancestor was an organism with the complexity of a prokaryote cell, and if viruses are essentially «escaped» material from complex genetic machinery, we cannot exclude the presence of viruses at a time close to the cenancestor. Knowing if the universal ancestor suffered viral infections or at least co-evolved with them requires further research, in the same way that many other questions in the study of the early evolution of life do.
Conclusions
There is a thread connecting our present with the first living beings, a thread formed by countless generations of ancestors and modified descendants. Generations that shape the universal tree of life on Earth once imagined by Darwin. Our inferences about the biology of the cenancestor are intimately linked to the structure of this universal tree and the evolutionary processes that shape it.
Arturo Becerra. Professor at the Science School of the National Autonomous University of Mexico (UNAM). He is the author of several scientific publications on the evolution of early life on Earth.
Luis Delaye. Professor at the Centre for Research and Advanced Studies at the National Polytechnic Institute (CINVESTAV), Irapuato Campus (Mexico). He is the author of several scientific publications on the evolution of bacteria.
REFERENCES
Agol, V. I. (2010). Which came first, the virus or the cell? Paleontological Journal, 44(7), 728–736. doi: 10.1134/S0031030110070038
Chatton, E. (1938). Titre et travaux scientifiques (1906–1937) de EdouardChatton. Sète: Sottano.
Darwin, Ch. (1859). On the origin of species by means of natural selection, or the preservation of favoured races in the struggle for life. Londres: John Murray.
Fitch, W. M., & Upper, K. (1987). The phylogeny of tRNA sequences provides evidence of ambiguity reduction in the origin of the genetic code. Cold Spring Harbor Symposia on Quantitative Biology, 52, 759–767. doi: 10.1101/SQB.1987.052.01.085
Koonin, E. V., & Dolja, V. V. (2013). A virocentric perspective on the evolution of life. Current Opinion in Virology, 3, 546–557. doi: 10.1016/j.coviro.2013.06.008
Lazcano, A., Fox, G. E., & Oró, J. (1992). Life before DNA: The origin and evolution of early Archean cells. In R. P. Mortlock (Ed.). The evolution of metabolic function (pp. 237–295). Boca Raton: CRC Press.
Margulis, L. (1975). Symbiotic theory of the origin of eukaryotic organelles: Criteria for proof. Symposia of the Society for Experimental Biology, 29, 21–38.
Mirkin, B. G., Fenner, T. I., Galperin, M. Y., & Koonin, E. V. (2003). Algorithms for computing parsimonious evolutionary scenarios for genome evolution, the last universal common ancestor and dominance of horizontal gene transfer in the evolution of prokaryotes. BMC Evolutionary Biology, 3(2). doi: 10.1186/1471-2148-3-2
Peretó, J., López-García, P., & Moreira, D. (2004). Ancestral lipid biosynthesis and early membrane evolution. Trends in Biochemical Sciences, 29(9), 469–477. doi: 10.1016/j.tibs.2004.07.002
Poole, A. M., Horinouchi, N., Catchpole, R. J., Si, D., Hibi, M., Tanaka, K., & Ogawa, J. (2014). The case for an early biological origin of DNA. Journal of Molecular Evolution, 79 (5–6), 204–212. doi: 10.1007/s00239-014-9656-6
Stanier, R. Y., & Van Niel, C. B. (1941). The main outlines of bacterial classification. Journal of Bacteriology, 42(4), 437–463.
Szathmáry, E. (2005). Life: In search of the simplest cell. Nature, 433, 469-470. doi: 10.1038/433469a
Whittaker, R. H. (1969). New concepts of kingdoms of organisms. Science, 163, 150–160. doi: 10.1126/science.163.3863.150
Woese, C. R., & Fox, G. E. (1977a). Phylogenetic structure of the prokaryotic domain: the primary kingdoms. Proceedings of the National Academy of Sciences, 74, 5088–5090. doi: 10.1073/pnas.74.11.5088
Woese, C. R., & Fox, G. E. (1977b). The concept of cellular evolution. Journal of Molecular Evolution, 10, 1–6. doi: 0.1007/BF01796132
Woese, C. R. (1998). The universal ancestor. Proceedings of the National Academy of Sciences, 95, 6854–6859.